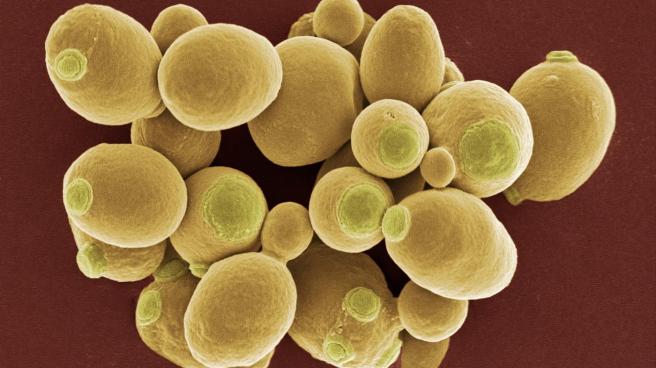
When I say the word ‘yeast’, you might think of making bread or beer.
One does not automatically think of Parkinson’s disease.
But yeast has actually been incredibly useful in enhancing our understanding of the genetics of Parkinson’s disease, and may well now provide us with novel treatments for the condition. In today’s post we will discuss how yeast research is leading the way for Parkinson’s disease.

Prof Susan Lindquist. Source: WallStreetJournal
It was with sadness that we heard about the passing of Prof Susan Lindquist in October last year. She was truly a pioneer in the field of molecular biology. In addition to advancing our understanding of gene functioning in degenerative diseases like Parkinson’s disease and Alzheimer’s, she also started a company, Yumanity, which is currently testing new drugs to tackle these conditions. Hopefully her legacy will have enormous impact for the millions of people around the world struggling with these conditions.
And that legacy all started with a bold (some even called it ‘crazy’) idea.
It involved yeast.
What is yeast?
Quite possibly the earliest domesticated species, yeast is a single-celled microorganism, traditionally classified as a member of the fungus kingdom. The evolutionary lineage of yeast dates back hundreds of millions of years old and there are at least 1500 species of yeast (Source: Wikipedia).
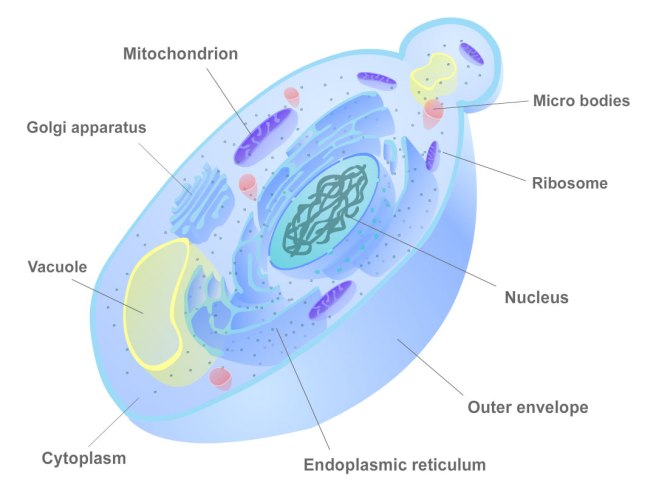
The cellular structure of yeast. Source: Biocourseware
More importantly, yeast is one of the most centrally important model organisms used in modern biological research, representing one of the most thoroughly researched organisms in the world.
We know more about the biology of yeast than we do about ourselves!
And this statement is made further evident as researchers use yeast to produce the world’s first synthetic organism (an organism for which the genome has been designed or engineered). By the end of 2017, the Synthetic Yeast 2.0 consortium plans to have produced a new form of yeast in which all 16 chromosomes will have been made in the lab (for more on this, read this STAT article).
Why do scientists like studying yeast?
The main reason is that yeast cells are very similar to human cells, but they grow a lot faster (human cells on average divide a rate of about once every 12 hours, while yeast cells divide every two hours). Yeast is similar to human cells in that it has all of the eukaryote structures, including a nucleus, cytoplasm, and mitochondria (eukaryote meaning a cell with a nucleus).
Yeast has played a fundamental role in many major scientific discoveries since the early 1900s. In 1907, German scientist Edward Buchner won the Nobel Prize in Chemistry for research involving yeast extract and fermentation. Ninety one years later (2006), Roger D. Kornberg won the same prize for his work on DNA transcription using yeast.
Yeast was the first eukaryote to have its genome (DNA) fully sequenced (in 1996). Yeast is literally leading the way in biological research.
So what has this got to do with Parkinson’s disease?
This is where Prof Susan Lindquist comes into the story.

Source: Youtube
In the early 2000s, she suggested the idea of using yeast to look at neurodegenerative conditions like Parkinson’s disease. It was a wild concept. Talking to the New York Times in 2007, she said “Even people in my laboratory thought we were crazy to try to study neurodegenerative diseases with a yeast cell. It’s not a neuron”.
But they persevered and in 2003 they published this research report in the journal Science:

Title: Yeast genes that enhance the toxicity of a mutant huntingtin fragment or alpha-synuclein.
Authors: Willingham S, Outeiro TF, DeVit MJ, Lindquist SL, Muchowski PJ.
Journal: Science. 2003; 302(5651):1769-72.
PMID: 14657499
In this study, the researchers conducted a genome-wide screens of mutant genes in yeast to identify genes that enhanced the toxic effects of the mutant huntingtin gene or alpha-synuclein protein. That is to say, they randomly mutated (made un-operational) just one gene per yeast cell, and these yeast cells either had the mutant huntingtin gene (which causes the neurodegenerative condition of Huntington’s disease) or too much alpha synuclein (which causes protein clumping in the yeast cells). Using this approach, they could determine which genes were responsible for increasing the negative effects of the mutant huntington gene or the alpha synuclein protein.
Of the 4850 yeast genes that the researcher mutated (and can we just point out that that is A LOT of work!!!), 52 were identified that exaggerated the affect of the mutant huntingtin gene and 86 increased sensitivity to alpha-synuclein (curiously, only one mutant gene resulted in increased sensitivity to both).
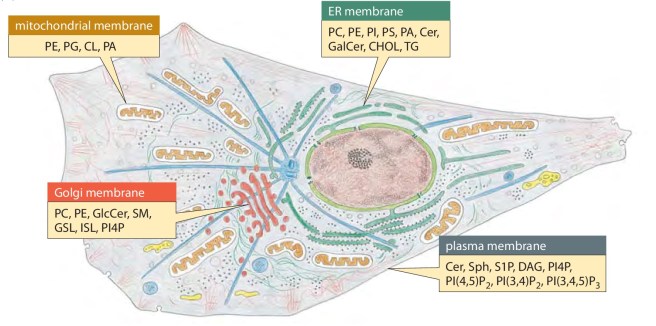
When they looked at the known functions of 86 genes that were increasing the sensitivity to alpha synuclein, the researchers found that most of them were involved in the processes of lipid metabolism (the synthesis and degradation of lipids) and vesicle-mediated transport (this occurs at the tips of neural branches – where alpha synuclein is located). Alpha synuclein is known to be involved with lipid metabolism (Click here and here for more on this). This reenforced the belief with the researchers that yeast could be used to assess disease relevant pathways – this study gave them the ‘proof of concept’.
In addition, the majority of the genes had human ‘orthologs’ (genes in different species that evolved from a common ancestral gene), meaning that the findings of yeast studies could potentially be translated to higher order creatures, like humans. The researchers concluded that they had found cell autonomous genes that are relevant to Parkinson’s disease.
With the publication of this work, people in Prof Lindquist’s lab were probably thinking the idea wasn’t so crazy anymore.
And this first publication led to many more, such as this report which was published in the same journal in 2006:

Title: Alpha-synuclein blocks ER-Golgi traffic and Rab1 rescues neuron loss in Parkinson’s models.
Authors: Cooper AA, Gitler AD, Cashikar A, Haynes CM, Hill KJ, Bhullar B, Liu K, Xu K, Strathearn KE, Liu F, Cao S, Caldwell KA, Caldwell GA, Marsischky G, Kolodner RD, Labaer J, Rochet JC, Bonini NM, Lindquist S.
Journal: Science. 2006 Jul 21;313(5785):324-8.
PMID: 16794039 (This article is OPEN ACCESS if you would like to read it)
In this study, the researchers doubled the amount of alpha synuclein that yeast cells produce and they observed that the cell stopped growing and started to die. They then looked at the earliest cellular events following theover production of alpha synuclein and they noticed that there was a blockage in the endoplasmic reticulum (ER)-to-Golgi vesicular trafficking.
Yes, I know what you are going to ask: What is ER-to-Golgi vesicular trafficking?
The endoplasmic reticulum (or ER) is a network of tubules connected to the nucleus. It is involved in the production of proteins and lipids, which are then transported to the Golgi apparatus which then modifies them, sorts them and and packs them into small bags called vesicles. These vesicles can then be taken to the cell membrane where the proteins are released to do their jobs.

The ER to Golgi pathway. Source: Welkescience
Now the fact that too much alpha synuclein was blocking this pathway was interesting, but the researchers wanted to go further. They conducted another genome-wide screens of genes in yeast to identify genes that could rescue this blockage – BUT this time, instead of mutating genes, the researchers randomly over produced one protein (and there was 3000 of them!!!) in each cell. Because the overproduction of alpha synuclein kills the yeast cells, all the researchers had to do was wait until the end of the experiment and determine which protein was over produced in the surviving cells.
And this led them to a protein called RAB1.
RAB1 is a protein that is critical to the transportation of proteins in cells. The researchers next tested the ability of RAB1 to rescue dopamine cells in Drosophila (flies), Caenorhabditis elegans (microscopic worms), and cell culture models of Parkinson’s disease and they found that it was able to rescue the cells in all three cases. While this result was very interesting, it also provided full validation of the approach that Prof Lindquist and her colleagues were taking using yeast cells to find new therapies for neurodegenerative conditions, like Parkinson’s disease.
But Prof Lindquist and her colleagues didn’t stop there.
Over the next decade numerous research reports were published taking advantage of this approach, including these two reports which appeared back-to-back in the journal Science:

Title: Identification and rescue of α-synuclein toxicity in Parkinson patient-derived neurons.
Authors: Chung CY, Khurana V, Auluck PK, Tardiff DF, Mazzulli JR, Soldner F, Baru V, Lou Y, Freyzon Y, Cho S, Mungenast AE, Muffat J, Mitalipova M, Pluth MD, Jui NT, Schüle B, Lippard SJ, Tsai LH, Krainc D, Buchwald SL, Jaenisch R, Lindquist S.
Journal: Science. 2013 Nov 22;342(6161):983-7.
PMID: 24158904
And:

Title: Yeast reveal a “druggable” Rsp5/Nedd4 network that ameliorates α-synuclein toxicity in neurons.
Authors: Tardiff DF, Jui NT, Khurana V, Tambe MA, Thompson ML, Chung CY, Kamadurai HB, Kim HT, Lancaster AK, Caldwell KA, Caldwell GA, Rochet JC, Buchwald SL, Lindquist S.
Journal: Science. 2013 Nov 22;342(6161):979-83.
PMID: 24158909
In these reports, Prof Lindquist and colleagues tested whether some of the proteins that had come up in their various screens could actually have positive benefits in human cells, specifically induced pluripotent stem (iPS) cells (which we have discussed before – click here to read that post). The researchers grew brains cells (neurons and glial cells) from the iPS cells derived from people who suffered from Parkinson’s disease with dementia. These cells exhibited a number of features that indicated that they were not healthy. From their yeast screens, the researchers identified a protein called Nedd4 that reversed the pathologic features in these neurons.
Nedd4 is an E3 ubiquitin-protein ligase, which is a protein involved in the removal of old or damaged proteins from a cell. It is part of the garbage disposal process. Importantly, Nedd4 has been shown to label alpha synuclein for disposal (Click here to read more about this) and it is also present in Lewy Bodies – the circular clusters of proteins present in the brains of people with Parkinson’s disease. Importantly, it is a ‘druggable’ target. Nedd4 has been considered a therapeutic target for cancer (Click here to read more on this).
More recently (and following the passing of Prof Lindquist) the group has published two research reports back-to-back in the journal Cell Systems:

Title: Genome-Scale Networks Link Neurodegenerative Disease Genes to α-Synuclein through Specific Molecular Pathways.
Authors: Khurana V, Peng J, Chung CY, Auluck PK, Fanning S, Tardiff DF, Bartels T, Koeva M, Eichhorn SW, Benyamini H, Lou Y, Nutter-Upham A, Baru V, Freyzon Y, Tuncbag N, Costanzo M, San Luis BJ, Schöndorf DC, Barrasa MI, Ehsani S, Sanjana N, Zhong Q, Gasser T, Bartel DP, Vidal M, Deleidi M, Boone C, Fraenkel E, Berger B, Lindquist S.
Journal: Cell Syst. 2017 Jan 25. pii: S2405-4712(16)30445-8.
PMID: 28131822 (This article is OPEN ACCESS if you would like to read it)
And:

Title: In Situ Peroxidase Labeling and Mass-Spectrometry Connects Alpha-Synuclein Directly to Endocytic Trafficking and mRNA Metabolism in Neurons.
Authors: Chung CY, Khurana V, Yi S, Sahni N, Loh KH, Auluck PK, Baru V, Udeshi ND, Freyzon Y, Carr SA, Hill DE, Vidal M, Ting AY, Lindquist S.
Journal: Cell Syst. 2017 Jan 25. pii: S2405-4712(17)30002-9.
PMID: 28131823
In these reports, Prof Lindquist and colleagues systematically mapped out molecular pathways underlying the toxic effects of alpha-synuclein. They applied their yeast derived 332 genes that impact alpha-synuclein toxicity, and linked them to multiple Parkinson’s-associated genes and druggable targets, using software called TransposeNet.

Source: Cell
The investigators then validated some of the connections in human iPS cells derived from people with Parkinson’s disease, confirming that some of the Parkinson’s disease-related genetic interactions observed in yeast are ‘conserved’ (that is maintained across evolution) to humans. And these findings fully vindicated Prof Lindquist’s ‘crazy’ idea of using yeast cells to investigate neurodegenerative disease.
In the second report, the researchers identified 225 proteins in close physical proximity to alpha-synuclein in neurons using a new technique (called ascorbate peroxidase (APEX) labeling – let’s just say it’s complicated, but if you’d like to read more about it, Click here). Many of those 225 proteins were well known to the researchers being involved with activities in vesicles and synaptic terminals, where alpha synuclein is often found. But the researchers also found microtubule-associated proteins (including tau) rubbing shoulders with alpha-synuclein, as well as proteins involved with mRNA binding, processing, and translation (which they were not expecting). Thus, not only has Prof Lindquist’s yeast model of Parkinson’s disease provided us with novel therapeutic targets, but also opened new avenues of research related to alpha synuclein functioning.
And there is now a ‘yeast’ biotech company?
It is called Yumanity.
Yumanity. Source: ScientificAmerican
Started in December 2014, with $45 million in funding, Yumanity is focused on determining new targets for neurodegenerative conditions in yeast cells, testing those new targets in human cells, and then moving towards clinical trials with the best candidates. To date, they have not announced any clinical trial candidates, but they working on compounds that are targeting the NEDD4 pathway in Parkinson disease (discussed above). We will be watching this company with great interest.
So what does it all mean?
Back in the early 2000s, Prof Lindquist and her team asked a simple but strange question:
Can we use our knowledge of yeast genetics to study neurodegeneration?
We now know that the answer is ‘yes’. The small single cell organism that most of us associate with baking and beer, shares enough genetics with us that we can use it as an assay for investigating molecular pathways involved with diseases of the brain. And in the not so distant future, this simple little organism may be providing us with new treatments and therapies for those diseases.
As we suggested at the start of this post, Prof Lindquist has left an amazing legacy. If Yumanity can move a new drug into clinical trials for Parkinson’s disease, it will only further strengthen that legacy.
Susan Lee Lindquist – June 5, 1949 to October 27, 2016
The banner for today’s post was sourced from NewEuropeans
 Yummy. Source: Healthline
Yummy. Source: Healthline Source: Visionpt
Source: Visionpt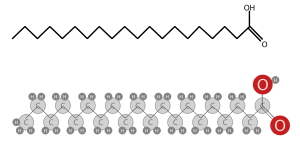 A chain of hydrocarbons ending in a carboxyl group (right). Source: Wikipedia
A chain of hydrocarbons ending in a carboxyl group (right). Source: Wikipedia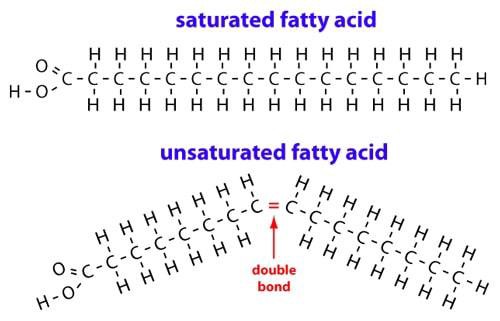 Saturated fatty acids vs unsaturated fatty acids. Source: Medium
Saturated fatty acids vs unsaturated fatty acids. Source: Medium Source: Medium
Source: Medium