|
Recently a Parkinson’s-associated research report was published that was the first of many to come. It involves the use of a genetic screening experiment that incorporates new technology called ‘CRISPR’. There is an absolute tidal wave of CRISPR-related Parkinson’s disease research coming down the pipe towards us, and it is important that the Parkinson’s community understands how this powerful technology works. In today’s post we will look at what the CRISPR technology is, how it works, what the new research report actually reported, and discuss how this technology can be used to tackle a condition like Parkinson’s. |
Me and my mother (and yes, the image is to scale). Source: Openclipart
My mother: Simon, what is all this new ‘crispy’ research for Parkinson’s I heard about on the news?
Me: Huh? (I was not really paying attention to the question. Terrible to ignore one’s mother I know, but what can I say – I am the black sheep of the family)
My mother: Yes, something about ‘crispy’ and Parkinson’s.
Me: Oh! You mean CRISPR. Yeah, it’s really cool stuff.
My mother: Ok, well, can you explain it all to me please, this ‘Crisper’ stuff?
Me: Absolutely.
CRISPR.101 (or CRISPR for beginners)
In almost every cell of your body, there is a nucleus.
It is the command centre for the cell – issuing orders and receiving information concerning everything going on inside and around the cell. The nucleus is also a storage bank for the genetic blueprint that provides most of the instructions for making a physical copy of you. Those grand plans are kept bundled up in 23 pairs of chromosomes, which are densely coiled strings of a molecule called Deoxyribonucleic acid (or DNA).
DNA’s place inside the cell. Source: Kids.Britannica
DNA is the ‘double helix’ stuff that biologists get excited about (Click here for the National Institute of Health’s website explanation of DNA). The information about you is stored in DNA using a code, made up of four chemical bases: adenine (A), cytosine (C), guanine (G), and thymine (T). These As, Cs, Gs and Ts are paired up and strung together in a huge number of pairings – approximately 3 billion pairs in a single cell in fact.
And when I say ‘pairs’, I mean that these letters pair off — A with T and C with G . An easy way to remember these pairings is Apples are in the Tree and Cars are in Garages:
Source: Pinterest
These pairings are called base pairs (the A, C, G & T being referred to as bases). Each base is also attached to a sugar molecule and a phosphate molecule. Together, these three components are called a nucleotide.
The AGTCs of DNA. Source: Knowgenetics
Understand that this simple pairing hold us together by holding our DNA together. They are critical to the normal biological functions of life – without them, there is no us.
Certain long segments of these pairings in the DNA provide the instructions for making a protein. These regions are called ‘Protein coding’ regions. These segments give rise to a molecule called Ribonucleic acid (or RNA) through a process called transcription. RNA is the intermediate that is used in the production of a protein – a process called translation.
Curiously, large portions of DNA is made up of regions that do not give rise to RNA, while other regions give rise to RNA, but no protein is made using that RNA. These region are called ‘Non-coding’ regions (the RNA that is produced has other functions in those situations – Click here to read more about this).
‘Protein coding’ or ‘Non-coding’. Source: Nature
Whether an RNA-producing region of DNA is a ‘Protein coding’ or ‘Non-coding’, these regions are all collectively referred to as genes. And it’s each of these genes that provide the instructions for keeping you alive.
BUT (and it’s a big BUT), with 3 billion different base pairs, you will understand that occasionally things may go a little awry. An individual pairing may have an error or a small section of DNA may be deleted, especially during periods when the DNA is being replicated (such as when cells divide). In many cases these errors do not have any effect. And in other cases, they may actually be beneficial (spontaneous errors are one of the foundations in the process of evolution).
Some times, however, these errors (also called genetic variants) can cause trouble.
Big trouble.
And these troublesome genetic variations come in different flavours: some can simply be a single mismatched base pair (also called a point-mutation or single nucleotide variant), while others are more complicated such as:

Lots of different types of genetic variations. Source: Nature
For the longest time…. well, basically all of human history, there has been no way to correct these variants. In fact, it’s only been in the last 60 years of homo sapiens existence that we have even become aware of them!
But the point is: Until now your DNA was set in stone – there was no way to correct it.
During the last 20-30 years, there have been efforts to insert functional bits of DNA into cells to replace the gene with an error in it. By inserting the healthy bit of DNA the cell can start to produce the required protein, and in most cases this restores normal functioning to the cell. But this has not altered the underlying fault in the DNA that required the insertion of functional DNA.
Then in 2013, this happened:
Title: Multiplex genome engineering using CRISPR/Cas systems.
Authors: Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F.
Journal: Science. 2013 Feb 15;339(6121):819-23.
PMID: 23287718 (This article is OPEN ACCESS if you would like to read it)
Not one research report, but two:
Title: RNA-guided human genome engineering via Cas9.
Authors: Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM.
Journal: Science. 2013 Feb 15;339(6121):823-6.
PMID: 23287722 (This article is OPEN ACCESS if you would like to read it)
These two research reports – published together – took the CRISPR-Cas9 system from two different bacteria Streptococcus thermophilus & Streptococcus pyogenes and inserted it into human cells. Using this system, the researchers then demonstrated very accurate editing of DNA, paving the way for the correction of troublesome genetic variants.
What does any of that actually mean?
Let’s go back to the beginning: 1987
It was a great year for some (for example, the Simpsons first appeared on TV as part of The Tracey Ullman Show)…
The Simpsons. Source: Wikipedia
…and a not-so-great year for others (think: the stock market crash in October).
It was also the year that some researchers at Osaka University in Japan noticed something really weird in the DNA of E.Coli bacteria: a set of 29-base pair repeats.
There they were. The same sequence of base pairs repeating over and over again in a short segment of the DNA. And they were separated by non-repeating, but similarly short sequences, that the researchers called ‘spacers’.
Repeats (in red) and spacers (in blue) in a region of bacterial DNA. Source: Rosalind
They published their finding in this report:

Title: Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product.
Authors: Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A.
Journal: J Bacteriol. 1987 Dec;169(12):5429-33.
PMID: 3316184 (This article is OPEN ACCESS if you would like to read it)
The observation left a lot of people in the biological research community scratching their heads. What none off these people realised at the time, however, was that this was the first report of a CRISPR locus.
What is a CRISPR locus?
Here we get to the heart of the matter.
A locus (plural: loci) in genetics is a fixed position or region of DNA. It is similar to a gene, but not necessarily functional – a locus is just a location.
And as I indicated above, Clustered Regularly Interspaced Short Palindromic Repeats (or CRISPR) are a series of repetitive sequences (exact repeats) interspaced by variable sequences (the spacers).
The big question surrounding these CRISPR regions was what the spacers actually did. What was their purpose?
The answer came in 2005, when three labs simultaneously and systematic analysed the spacer sequences and discovered that they were ‘extrachromosomal‘ – in other words, originating from a different species DNA and probably not really supposed to be there (Click here, here and here to see those three research reports).
They also noted that the spacer DNA segments were exact copies of sections of DNA from phage.
What are phage?
Phage are viruses that infects and replicate within a bacteria. Bacteria really don’t like them.
Phage life cycle. Source: Medium
The finding of phage DNA in the CRIPSR region caused the researchers to start speculating that maybe these CRISPR regions were playing a role in the immune defence system of the bacteria.
And this idea was then tested:
Title: CRISPR provides acquired resistance against viruses in prokaryotes.
Authors: Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P.
Journal: Science. 2007 Mar 23;315(5819):1709-12.
PMID: 17379808
In this study, the researchers took phage-resistant bacteria and infected them with a specific type of phage that the bacteria had never been exposed to in the past. They found that after the phage infections, these bacteria integrated small segments of the phage DNA (new spacers derived from phage DNA) into some of the CRISPR regions.
They also found that removal or addition of a particular ‘spacer’ modified the phage-resistance of a bacteria. By adding a spacer of phage DNA to the bacteria’s CRISPR regions, the researchers could make the bacteria resistant to that phage. And by removing a particular spacer of phage DNA from the bacteria’s CRISPR regions, the researchers could make the bacteria more vulnerable to that phage.
The question then became: how are these regions of bacterial DNA doing this? How are these ‘spacers’ having this effect?
Do we know?
Yes, we do. Over the last 10 years it has been thoroughly researched and we now understand the process very well.
During the infection of a bacteria, a phage will inject its DNA into the cell. This DNA will be recognised as foreign DNA by two proteins (called Cas1 and Cas2), and they will chop it up into small pieces which will then be inserted into a CRISPR locus in the bacterial DNA. And once it is embedded in the DNA, these spacers can be passed on to any further cells, if that bacteria decides to divide.
CRISPR regions of the DNA are regularly being transcribed (that is the process of producing RNA). The RNA for each spacer region is called crRNA (CRISPR-RNA), and it is basically a small RNA molecule whose sequence matches a region of phage DNA. This crRNA then attaches to another piece of RNA to become what is called a tracrRNA.
This tracrRNA then joins the Cas9 protein and together they start wandering around the bacteria looking for sections of DNA to check for that particular crRNA.
How CRISPR works in bacteria. Source: Sciencedirect
Hang on a second. What is Cas9?
Cas9 (or CRISPR associated protein 9) is the important piece of the puzzle. It is the protein that does the magic.
It is an endonuclease.
What is an endonuclease?
As I suggested above, replication of DNA is never perfect and there are occasionally errors. Millions of years of evolution has given rise to a very sophisticated, but extremely efficient system of DNA monitoring within cells. Many different proteins are involved in this process, and endonucleases play an important role.
An endonucleases is an enzyme that binds to DNA and cuts it. Importantly, it functions in the middle of the chain of DNA, while another class of enzyme (called exonucleases) operate from the end of the chains.
So Cas9 is an enzyme that can cut DNA in the middle of the chain of DNA?
Exactly. And it does this in a very targeted manner, being guided by pieces of RNA (the crRNA mentioned above) that are produced from the CRISPR regions of bacterial DNA. The crRNA is now often referred to as the ‘guide RNA’.
Cas9 protein interacting with DNA. Source: Stackexchange
And if a particular Cas9 enzyme joins forces with a tracrRNA and then finds a matching piece of phage DNA floating around inside the bacteria, then the bacteria will then know that this particular type of phage has infected it and the phage DNA should be disposed of ASAP. I should add that that this is not a conscious process for the bacteria – it is simply an acquired, automatic immune response that helps protect the bacteria.
Interestingly, only 40% of sequenced bacteria and 90% of archaea (single cell organisms that are separate to bacteria) have this CRISPR-Cas9 system. How the rest of the bacterial world are protecting themselves from phage and other nasties is an area of enormous research efforts.
Sounds interesting, but how is this being applied to humans?
This is where the story gets really interesting, and not just from the stand point of incredibly powerful technology.
You see, there is a really big legal battle behind the CRISPR technology (and I suspect that the lawyers will be the only winners at the end of it all – Click here and more recently here to read about this legal argument). The arguments are so furious that anyone writing about this next chapter of CRISPR history has to be careful, because they are sure to upset someone about the actual order of things.
Basically several different research organisations managed to insert CRISPR-Cas9 into other organisms (including mammalian cells) AND demonstrate targeted DNA editing. And they all did it at roughly the same time. Given the potential applications of this genome-editing technology, everybody wants to be seen as doing it first for the sake of being the inventor and thus reaping the financial rewards (hence the legal arguments).
From the stand point of human cells, we are back to the two research reports (mentioned above) from 2013:

Title: Multiplex genome engineering using CRISPR/Cas systems.
Authors: Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F.
Journal: Science. 2013 Feb 15;339(6121):819-23.
PMID: 23287718 (This article is OPEN ACCESS if you would like to read it)

Title: RNA-guided human genome engineering via Cas9.
Authors: Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM.
Journal: Science. 2013 Feb 15;339(6121):823-6.
PMID: 23287722 (This article is OPEN ACCESS if you would like to read it)
In these two research reports, the investigators took a CRISPR region from two different bacteria Streptococcus thermophilus and Streptococcus pyogenes, and they then designed and inserted spacers/crRNA for a particular human gene into that CRISPR region. Next, they inserted this CRISPR region and the Cas9 protein into the human cells and watched to see what would happen to that particular gene.
The results were very clear – they had demonstrated very accurate editing of DNA.
And it not fake news to suggest that these findings literally changed the world.
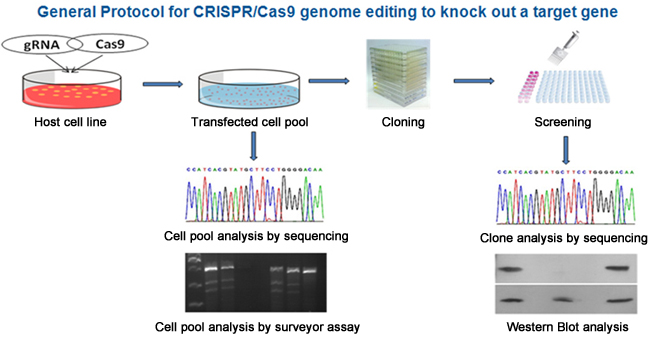
Since 2013, CRISPR-Cas9 has become a topic that is discussed on a daily basis in laboratories all around the world. Everyone in biomedical research is now trying to incorporate this technology into some aspect of their work. And the approach has become much simplified: now researchers simply insert into a cell the Cas9 protein and specific guideRNAs for the target gene that they want to mutate and then they let the cell do the rest.
CRISPR experiment for removing (knocking-out) the function of a gene. Source: Genscript
And the CRISPR-Cas9 technology has also been re-engineered in different ways to allow alternative functions to be introduced. For example, the Cas9 protein has been re-engineered so that rather than cutting DNA, the Cas9 protein now activates that region of DNA where it is directed to be the guideRNA (Click here to read more about this).
I appreciate that all of this biology is rather complex and may be confusing, and if none of what I have written above about CRISPR makes any sense, please watch this short video in which Feng Zhang explains how CRISPR works:
(Feng is one of the leading researchers looking at CRISPR – and a likely Nobel prize winner in the near future)
It is impossible to understate what a revolution in biological science CRISPR-Cas9 has generated. The ability to finely edit and alter DNA – cutting out individual bases and introducing something new – has changed the way researchers design their studies. And there is an endless list of possible applications in medical research.
Take, for example, AIDS (acquired immune deficiency syndrome) research.
Currently people infected with the human immunodeficiency virus (HIV) must use a complex concoction of anti-retroviral medications to suppress the virus and stop it from replicating and developing into full blown AIDS (which would result in the immune system being unable to fight infections).
Recently, however, researchers have effectively ‘cured’ mice with the disease, but introducing the CRISPR-Cas9 technology into the blood cells of the mice:
Title: In Vivo Excision of HIV-1 Provirus by saCas9 and Multiplex Single-Guide RNAs in Animal Models.
Authors: Yin C, Zhang T, Qu X, Zhang Y, Putatunda R, Xiao X, Li F, Xiao W, Zhao H, Dai S, Qin X, Mo X, Young WB, Khalili K, Hu W.
Journal: Mol Ther. 2017 May 3;25(5):1168-1186.
PMID: 28366764
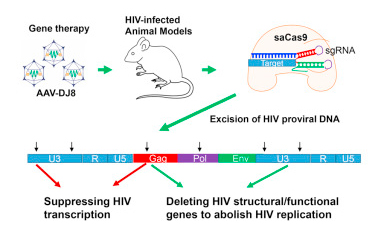
In this study, the researchers designed guide RNAs for sections of the HIV DNA. After infecting a cell, the HIV virus will embed the instructions for replicating itself in the DNA of the infected cell. By introducing the Cas9 protein and guideRNAs for sections of the HIV DNA, the investigators were able to lower levels of HIV by up to 95% in live mice. The virus could infect the blood cells, but if the CRISPR-Cas9 technology was present in the cell, the virus wouldn’t be able to replicate itself. Over time, the blood cells with the CRISPR-Cas9 technology will have a competitive advantage over the blood cells that do not, and these cells will restrict the ability of HIV to progress to full blown AIDS.
Source: Cell
The researchers behind this study suggest that “The next stage would be to repeat the study in primates, a more suitable animal model where HIV infection induces disease, in order to further demonstrate elimination of HIV-1 DNA in latently infected T cells and other sanctuary sites for HIV-1, including brain cells. Our eventual goal is a clinical trial in human patients” (Source).
It is a remarkable application of the CRISPR-Cas9 technology that has very real world implications.
I really don’t think I’m exaggerating when I write that the potential of the technology is truly limitless (Click here to read various OPEN ACCESS articles in Cell press that highlight this fact). The only factor restricting us is our imagination.
But with such ability comes great responsibility.
There is already a great deal of idiotic talk of designer babies, but such possibilities are still a long way off in the future (Click here, here, here and here to read interesting news articles on this topic).
The reason for the delay in designing ‘perfect little humans’ is that the gap between our ability to manipulate genes and our understanding of what genes actually do is vast. We simply don’t know enough about what each gene does in order to be confident in starting to editing things laissez faire.
Take, for example, the simple case of red hair: Say a family want their next unborn child to have red hair. The requirements are very straightforward: alter certain regions of the MC1R gene and – hey presto – you have a ginger head baby.
A wannabe redhead…cute cheeks though. Source: HuffingtonPost
But our understanding of the function of the MC1R gene is still very basic and we do not know the full impact of such a manipulation. Remember, redhead people (those with the MC1R genetic variant) are more likely to develop melanoma and possibly Parkinson’s disease (though we are not sure why).
Thus, we must proceed very carefully with how we handle this powerful new CRISPR technology.
Having said all of that, the pace of CRISPR-Cas9 research is breath taking!
Every few days a remarkable new CRISPR-based tool is being reported (just last week a modified version of CRISPR-Cas9 was announced that is able to change a single base (one of those C, A, T, or G letters) at a time without actually making any breaks in the DNA’s structure (Click here to read more about this)!
So what is the new CRISPR-related Parkinson’s research?
Title: A CRISPR screen identifies a pathway required for paraquat-induced cell death.
Authors: Reczek CR, Birsoy K, Kong H, Martínez-Reyes I, Wang T, Gao P, Sabatini DM, Chandel NS.
Journal: Nat Chem Biol. 2017 Oct 23.
PMID: 29058724
In this study, the researcher grew human leukemia cells that have the Cas9 protein inserted into them in cell culture. They then infected the cells with a large pool of viruses – the viruses were identical except for one critical difference: each virus had a single guideRNA that targets just one of 3,000 different genes associated with cell metabolism (involving basically all of the chemical changes that are involved in energy and basic functioning in a cell).
Thus, each cell in the culture of millions of cell would have just one of the 3000 genes randomly mutated. This approach allowed the investigators to determine in large screening studies which genes are important for a particular function.
And the function that the investigators in this study wanted to assess was which genes are involved in the handling of a specific toxin.
They cultured the cells in the presence of the herbicide ‘paraquat’ – a common weedkiller that has been associated with Parkinson’s (Click here to read an OPEN ACCESS review of this). And after exposing the cell cultures to paraquat for 10 days, the investigators discovered three genes (POR, ATP7A, and SLC45A4) are required for paraquat-induced cell death. When these genes are mutated, the toxic effects of paraquat are significantly reduced.
POR was particularly potent.
POR (cytochrome P450 oxidoreductase) is an enzyme that has previously implicated in the generation of oxidative stress in cells, and the researchers in this study demonstrated that it is required for this process. Thus, this research may have identified a novel target for future therapeutic drugs.
And this CRISPR-based screening approach has been used in Parkinson’s research in other ways as well, such as identifying genes that protect against the toxic version of the Parkinson’s associated protein, alpha synuclein:
Title: Randomized CRISPR-Cas Transcriptional Perturbation Screening Reveals Protective Genes against Alpha-Synuclein Toxicity.
Authors: Chen YC, Farzadfard F, Gharaei N, Chen WCW, Cao J, Lu TK.
Journal: Mol Cell. 2017 Oct 5;68(1):247-257.
PMID: 28985507
In this study, the researchers used a yeast model of Parkinson’s.
Yes, yeast.
Yeast cells. Source: NewEuropeans
We have previously discussed the idea of using yeast as a model of Parkinson’s (Click here to read that post). While the yeast cells do not exhibit many of the physical (motor) features of Parkinson’s, they do share many of the same genes as us and this commonality allows us to use them as a cellular model for exploring the genetic interactions of the condition. In this study, the investigators used yeast cells that have been genetically engineered to produce A LOT of the Parkinson’s associated protein, alpha synuclein.
The researchers used these yeast cells to perform a CRISPR-based screen of genes that protect the cells against the toxic version of alpha synuclein. That experiment identified two guideRNAs (designated as gRNA 6-3 and gRNA 9-1) that had a powerful protective effect – that is, by mutating the genes associated with those guideRNA, the researchers could infer protection on the cells against the effects of the toxic alpha synuclein.
The more powerful of the two guideRNAs was gRNA 9-1…but there was a slight problem: the researchers could not clearly identify where in the DNA this guideRNA was targeting.
So they could not determine which specific gene was responsible for the powerful effect!
This did not stop the researchers though. They next looked at what the consequences of gRNA 9-1 targeting were. Specifically they were interested in which genes were being affected by the actions of the gRNA 9-1 targeted mutation. They identified 114 genes that had more than a 2-fold changes in their level of activity.
And the top gene: DJ-1
This is a really interesting result because DJ-1 – also known as PARK7 – is a gene with a long association with Parkinson’s (Click here to read a previous post on DJ-1). By over-producing DJ-1 in the yeast cells with high levels of alpha synuclein, the researchers found that they could reduce levels of cell death. The investigators also demonstrated this result in human cells. This DJ-1 effect on alpha synuclein has also been demonstrated by previous groups (Click here and here to read about those studies).
Interestingly, the investigator’s screening experiment also identified two additional genes (TNX and TIMM9) which both play important roles in reducing oxidative stress in cells. But by increasing the activity of these genes individually, the researchers failed to rescue the toxic effects on alpha synuclein. It was only when they increased activity levels of both genes that they saw a complete rescue of the cell loss associated with alpha synuclein. The researchers are now looking for activators of these genes which could potentially be turned into therapeutic treatments for Parkinson’s.
Very interesting. But this CRISPR technology is so new, I guessing no clinical trials have been conducted yet?
You would be wrong in thinking that.
In fact, in June of last year (2016 – just 3 years after CRISPR was put into human cells), US researchers got the green light from an advisory committee at the US National Institutes of Health (NIH) for a proposal to use CRISPR–Cas9 to treat cancer. And exactly one year ago (on 28th October, 2016), Chinese researchers started a clinical trial that involves CRISPR-modified cells being injected into patients with aggressive lung cancer (Click here to read more about this).
And last month, the European Medicines Agency (EMA) granted orphan medicinal product status to Editas Medicine Inc for their pre-clinical, CRISPR-based medicinal product EDIT-101 (Click here to read more).
The company hopes to treat Leber Congenital Amaurosis type 10 (a genetic disorder which is the most common cause of inherited childhood blindness; 3 per 100,000 live births worldwide). Editas Medicine plans to submit an Investigational New Drug (IND) application for EDIT-101 in mid-2018.
In addition to all of this activity, all of the big biotech and pharmaceutical companies are building up their CRISPR-Cas9 screening resources. It is a modern day gold-rush and the impact of it will hopefully have major benefits for all of us.
And again, remember that all of this is happening just 3-4 years after the CRISPR-Cas9 mechanism was used in human cells for the first time. Again – it goes without saying – this field is moving with breath-taking speed!
What about clinical trials for Parkinson’s disease?
So while there are genetic variants associated with Parkinson’s (see our Genetics of Parkinson’s page for more on this), it is unlikely that we will be seeing any CRISPR-correcting clinical trials of those genetic alterations in the near future.
The problem is ‘delivery’.
How does one deliver the CRISPR-Cas9 technology to every cell in the body? Or even just every cell in the brain? Delivery has alway been a major issue in medicine (it is one of the main reasons drugs have side effects). Some readers may point towards a recent post in which I discussed a new gene therapy virus that only infects brain cells when injected into the body (Click here to read that post). While this approach may seem ideal for the task of correcting a PARK gene variant in the brain, the Adeno-associated virus (AAV) that is used in those experiment can not carry enough DNA to deliver all of the CRISPR-Cas9 technology required for the task.
The news is not all bad though.
There are a lot of CRISPR-based screening experiments currently ongoing that will identify a lot of genes which could be beneficial for Parkinson’s. Those studies will hopefully lead to novel therapies – or even better point towards clinically available drugs – that can be tested in the clinic. And given the pace of all the CRISPR research, I doubt that we will have to wait more than 6 months before those screening experiments start to result in research reports.
What does it all mean?
So summing up (wow, long post!): You may not be aware of this, but we are currently in the midst of a 21st century gold rush.
DNA borrowed from bacteria has revolutionised the way biological research is being conducted. New tools based on this CRISPR technology are speeding up the progress of biomedical research, and leading to the identification of novel therapeutic targets and new ways of treating medical conditions.
This post is rather long, but it is important for folks in the Parkinson’s community to be aware of this new CRISPR-Cas9 technology and how it works. It is going to have major implications for Parkinson’s research moving ahead and perhaps it may even be involved with clinical trials in the not-so-distant future (I am happy to be wrong in my prediction mentioned above). It is really very exciting to watch all of the new discoveries being made using this technology and I look forward to reporting the new findings as they get published. This post was also put here as a primer, that future CRISPR-related posts can be pointed towards.
Lots of ‘CRISPY’ Parkinson’s research coming this way!
The banner for today’s post was sourced from EIC




















Hi Simon
Very many thanks for taking the time to prepare this post, which is so important to our understanding of the way forward in Parkinson’s research. I really appreciate the way in which you make scientific papers so readily understandable for the wider community, and I love your style of writing! Please keep up the good work – it is invaluable.
Best wishes
Ken
LikeLike
Hi Ken,
Thanks for the kind words – glad you liked the post. This CRISPR stuff is fascinating and I could have gone on a lot longer, but I feared that I might lose a few readers if I didn’t stop where I did. Glad it all made sense to someone.
Kind regards,
Simon
LikeLike
I cannot tell you how grateful I am to have followed this blog for the past few months since stumbling upon it. My husband and I care for my mother who has had Parkinson’s Disease for almost 20 years. At the same time, we are parenting young children. My father is also showing symptoms and I have fears that I will also be diagnosed at some point down the road due to welding that my father did in the household when I was a child. Your explanations and summaries of the current research is a bright spot. I don’t know what else to say except thank you from the bottom of my heart for taking the time to do this blog!
LikeLike
I, too, find these posts leave me feeling positive even if a bit overwhelmed with good info. Thanks again
LikeLike
Awesome! CRISPR sounds like the biological equivalent of going from the stone age to the iron age. I wonder what the next breakthrough biological tool might look like?
LikeLike
There is a new cas9 enzime whose genectic code can fit inside the AAV virus. Its called mini cas9 or saCas9.
LikeLike
So it can be used in gene therapy to replace a faulty gene.
LikeLike
Hi Gui,
Thanks for your comment. An AAV with the SaCas9 and gRNAs is already available (https://www.addgene.org/61591/). I have some colleagues who are working with one of them in the shop. And I’m sure that researchers somewhere in the world are already trying to squeeze SaCas9 into the new AAV-PHP viruses which allow for non-invasive gene delivery (in mice at least – https://scienceofparkinsons.com/2017/12/10/monkeys/). Given the pace of everything at the moment, I’d be surprised if we don’t see a lot of research involving this technology in 2018.
Kind regards,
Simon
LikeLike
Simon, great article. But now you have a responsibility, you have to keep us updated. My mom has Parkinson’s Plus and I feel crispr is the holy grail for a cure or almost a cure. Please keep us posted on any breakthroughs.
LikeLike
Hi Mic,
Thanks for your comment – glad you liked the post. Sorry to hear about your mothers diagnosis, I hope she is responding well to the treatments.
CRISPR has incredible potential. And just here in Cambridge (UK) I am aware of numerous Parkinson’s-related CRISPR projects currently being conducted. There will be a lot of data related to this coming in 2018. It is going to be a big year! And I’ll be happy to try and make some sense of it all here on the SoPD. Stay tuned.
Kind regards,
Simon
LikeLike