|
Last week, as everyone was preparing for Christmas celebrations, researchers at the pharmaceutic company Novartis published new research on a gene that is involved with Parkinson’s, called PARKIN (or PARK2). They used a new gene editing technology – called CRISPR – to conduct a large screening study to identify proteins that are involved with the activation of PARKIN. In today’s post we will look at what PARKIN does, review the research report, and discuss how these results could be very beneficial for the Parkinson’s community. |
Source: Novartis
As many people within the Parkinson’s community will be aware, 2017 represented the 200th anniversary of the first report of Parkinson’s disease by James Parkinson.
It also the 20th anniversary of the discovery of first genetic mutation (or variant) that increases the risk of developing Parkinson’s. That genetic variation occurs in a region of DNA (a gene) called ‘alpha synuclein’. Yes, that same alpha synuclein that seems to play such a critical role in Parkinson’s (Click here to read more about the 20th anniversary).
In 2018, we will be observing the 20th anniversary of the second genetic variation associated with Parkinson.
That gene is called PARKIN:
Title: Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism.
Authors: Kitada T, Asakawa S, Hattori N, Matsumine H, Yamamura Y, Minoshima S, Yokochi M, Mizuno Y, Shimizu N
Journal: Nature. 1998 Apr 9; 392(6676):605-8
PMID: 9560156
In 1998, Japanese researchers published this report based on 5 individuals from 4 Japanese families who were affected by juvenile-onset Parkinson’s. In family 1, the affected individual was a female, 43 years old, born of first-cousin parents, and her two younger brothers are healthy. Her condition was diagnosed in her teens and it had then progressed very slowly afterwards. Her response to L-dopa was very positive, but L-dopa-induced dyskinesia were frequent. In family 2-4, affected individuals (born to unrelated parents) exhibited very similar clinical features to the subject in family 1. The age of onset was between 18 to 27 years of age.
Using previous research and various techniques the investigators were able to isolate genetic variations that were shared between the 5 affected individuals. They ultimately narrowed down their search to a section of DNA containing 2,960 base pairs, which encoded a protein of 465 amino acids.
They decided to call that protein PARKIN.
PARKIN Protein. Source: Wikipedia
How much of Parkinson’s is genetic?
Approximately 15% of people with Parkinson disease have a family history of the condition – a grandfather, an aunt or cousin.
At present, about 10-20% of Parkinson’s disease cases can be accounted for by genetic variations that infer a higher risk of developing the condition. In people with ‘juvenile-onset’ (diagnosed under the age 20) or ‘early-onset’ Parkinson’s disease (diagnosed under the age 40), genetic variations can account for the majority of cases, while in later onset cases (>40 years of age) the frequency of genetic variations is more variable.
For a very good OPEN ACCESS review of the genetics of Parkinson’s disease – click here.
There are definitely regions of DNA in which genetic variations can increase one’s risk of developing Parkinson’s disease.
These regions are referred to as PARK genes.
What are PARK genes?
We currently know of 23 regions of DNA that contain mutations that have causal-associations with Parkinson’s. These regions have been given the name of PARK genes. The region does not always refer to a particular gene, for example in the case of our old friend alpha synuclein, there are two PARK gene regions within the stretch of DNA that we call alpha synuclein. So please don’t think of each PARK genes as one particular gene.
In addition, there can be multiple genetic variations within a PARK gene that can infer risk of developing Parkinson’s disease. The increased risk is not always the result of one particular mutation within a PARK gene region.
PARKIN is referred to as PARK2.
Where in the PARKIN gene is the mutation located?
So this is where things get a bit more complicated.
There isn’t one single genetic variation associated with Parkinson’s. In fact, there are currently 10 common mutations in the PARKIN gene that can give rise to early-onset Parkinson’s disease. PARKIN has also been associated with different types of cancer – there are 13 cancer-related variants. Importantly, only two of the Parkinson’s related variants are associated with an increased risk of cancer (they are R24P and R275W – red+black arrow heads in the image below).

Comparing PARK2 Cancer and PD associated mutations. Source: Nature
Thus it is important to know exactly where your mutation is, if in fact you have one. And if you have the R24P and R275W mutation, it doesn’t necessarily mean that you are going to get a particular cancer. It just means that you are slightly more predisposed to it. So it is good to be aware of this and to have regular check ups.
PARKIN protein has numerous functions within a cell. But it is primarily considered an E3 ubiquitin ligase – this is one of several enzymes involved in a process that is called ubiquitination.
What is ubiquitination?
Ubiquitin is a small protein that, as the name suggests, is ubiquitous in all cells, and it affects all aspects of cell biology. Enzymes like PARKIN attach ubiquitin to particular proteins (the protein goes through the process of ubiquitination) and this has different effects on that protein. Ubiquitination can cause a protein to be moved to a different cellular location, or affect the level of activity for that protein, and it can even promote or prevent certain interactions between different proteins.
But the most studied process associated with ubiquitination is the marking of particular proteins for disposal. A protein labelled with ubiquitin in a particular way will be sent off for disposal and be broken down in a structure called the Proteasome:
PARKIN is involved in the Ubiquitin-E3 step. Source: 2bscientific
For a more thorough explanation of the Ubiquitin Proteasome pathway, please watch this video provided by the Scottish Enterprise:
OK, but you said that PARKIN protein has numerous functions. What else does it do?
It is also involved with the process of disposing of old or dysfunctional mitochondria.
Mitochondria – you may recall from previous SoPD posts – are the power stations of each cell. They help to keep the lights on. Without them, the party is over and the cell dies.

Mitochondria and their location in the cell. Source: NCBI
You may remember from high school biology class that mitochondria are tiny bean-shaped objects within the cell. They convert nutrients from food into Adenosine Triphosphate (or ATP). ATP is the fuel which cells run on. Given their critical role in energy supply, mitochondria are plentiful (some cells have thousands) and highly organised within the cell, being moved around to wherever they are needed.
Like you, me and all other things in life, however, mitochondria have a use-by date.
As mitochondria get old and worn out (or damaged) with time, the cell will recycle them via a process called mitophagy (a blending of the words mitochondria and autophagy – the waste disposal system of each cell).
PARKIN plays an important role in mitophagy.
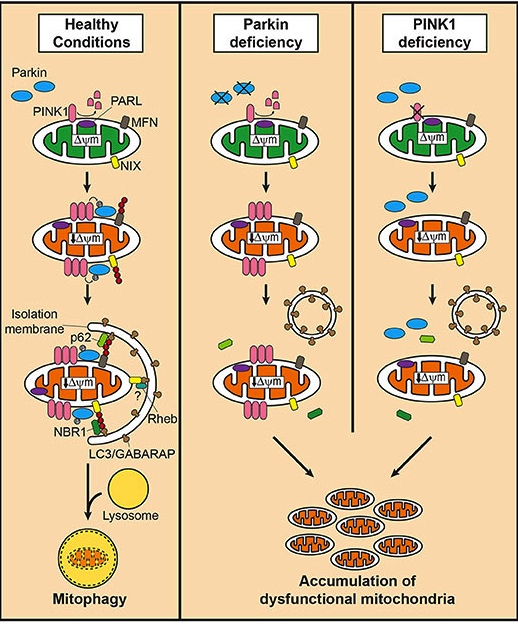
What does PARKIN do in Mitophagy?
PARKIN interacts with another Parkinson’s-associated protein called PINK1.
PINK1 acts like a kind of handle on the surface of mitochondria. In normal, healthy mitochondria, the PINK1 protein attaches to the surface of mitochondria and it is slowly absorbed until it completely disappears from the surface and is degraded. In unhealthy mitochondria, however, this process is inhibited and PINK1 starts to accumulate on the outer surface of the mitochondria.
Lots of handles poking out of the surface of the mitochondria.
Now, if PINK1 a handle, then PARKIN is the flag that likes to hold onto the PINK1 handle. While exposed on the surface of mitochondria PINK1 starts grabbing the PARKIN protein. This pairing (along with a bit of ubiquitination) is a signal to the cell that this particular mitochondrion (singular) is not healthy and needs to be removed.

Pink1 and Parkin in normal (right) and unhealthy (left) situations. Source: Hindawi
In the absence of normal PINK1 or PARKIN proteins, there is no handle-flag system and sick/damaged mitochondria start to pile up. They are not disposed of appropriately and as a result the cell getting sick and ultimately dying.
Mitophagy. Source: Frontiersin
In addition to mitophagy, PARKIN may also act as a tumor suppressor protein, which means it prevents cells from growing and dividing too rapidly or in an uncontrolled way (Click here to read more about this). And PARKIN may also regulate the supply and release of tiny sacks of neurotransmitter (called synaptic vesicles) from nerve cells (Click here to read more about this).
How is the PARKIN mutation caused?
The juvenile-onset form of PARK2-associated Parkinson’s is caused by an autosomal recessive mutation – meaning that a copy of the mutation has to be provided by both the parents in order for a condition to develop.
Autosomal recessive genetic transfer. Source: Wikipedia
OK, so what is this research that Novartis has published recently?
It involves a new gene editing technology called CRISPR.
What is CRISPR?
I have previously written a long post explaining the science behind CRISPR (Click here to read that post).
Briefly, Clustered Regularly Interspaced Short Palindromic Repeats (or CRISPR) are a series of repetitive sequences (exact repeats) of DNA that were found in particular bacteria, which form a frontline defensive mechanism against infection from nasty viruses.
During the infection of a bacteria, a phage (a virus that infects bacteria) will inject its DNA into the cell. This DNA will be recognised as foreign DNA by two proteins (called Cas1 and Cas2), and they will chop it up into small pieces which will then be inserted into the bacterial DNA (a region referred to as a CRISPR locus). And once it is embedded in the DNA, this CRISPR locus can be passed on to any further cells, if that bacteria decides to divide.
CRISPR regions of the DNA are regularly being transcribed (that is the process of producing RNA). The RNA for each spacer region is called crRNA (CRISPR-RNA), and it is basically a small RNA molecule whose sequence matches a region of phage DNA. This crRNA then attaches to another piece of RNA to become what is called a tracrRNA.
This tracrRNA then joins the Cas9 protein and together they start wandering around the bacteria looking for sections of DNA to check for that particular crRNA. Like a guard wandering around on patrol duty, looking for signs of trouble at a prison.
How CRISPR works in bacteria. Source: Sciencedirect
Hang on a second. What is Cas9?
Cas9 (or CRISPR associated protein 9) is the important piece of the puzzle. It is the protein that does the magic.
It is an endonuclease.
What is an endonuclease?
As I suggested above, replication of DNA is never perfect and there are occasionally errors. Millions of years of evolution has given rise to a very sophisticated, but extremely efficient system of DNA monitoring within cells. Many different proteins are involved in this process, and endonucleases play an important role.
An endonucleases is an enzyme that binds to DNA and cuts it. Importantly, it functions in the middle of the chain of DNA, while another class of enzyme (called exonucleases) operate from the end of the chains.
So Cas9 is an enzyme that can cut DNA in the middle of the chain of DNA?
Exactly. And it does this in a very targeted manner, being guided by pieces of RNA (the crRNA mentioned above) that are produced from the CRISPR regions of bacterial DNA. The crRNA is now often referred to as the ‘guide RNA’.
Cas9 protein interacting with DNA. Source: Stackexchange
And if a particular Cas9 enzyme joins forces with a tracrRNA and then finds a matching piece of phage DNA floating around inside the bacteria, then the bacteria will then know that this particular type of phage has infected it and the phage DNA should be disposed of ASAP. I should add that that this is not a conscious process for the bacteria – it is simply an acquired, automatic immune response that helps protect the bacteria.
Interestingly, only 40% of sequenced bacteria and 90% of archaea (single cell organisms that are separate to bacteria) have this CRISPR-Cas9 system. How the rest of the bacterial world are protecting themselves from phage and other nasties is an area of enormous research efforts.
Sounds interesting, but how is this being applied to humans?
Way back in the old times (think 2013) two research reports reported successful use of the CRISPR-Cas9 system in human cells:

Title: Multiplex genome engineering using CRISPR/Cas systems.
Authors: Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F.
Journal: Science. 2013 Feb 15;339(6121):819-23.
PMID: 23287718 (This article is OPEN ACCESS if you would like to read it)

Title: RNA-guided human genome engineering via Cas9.
Authors: Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM.
Journal: Science. 2013 Feb 15;339(6121):823-6.
PMID: 23287722 (This article is OPEN ACCESS if you would like to read it)
In these two research reports, the investigators took a CRISPR region from two different bacteria Streptococcus thermophilus and Streptococcus pyogenes, and they then designed and inserted spacers/crRNA for a particular human gene into that CRISPR region. Next, they inserted this CRISPR region and the Cas9 protein into the human cells and watched to see what would happen to that particular gene.
The results were very clear – they had demonstrated very accurate editing of human DNA.
And it not fake news to suggest that these findings literally changed the world.
Since 2013, CRISPR-Cas9 has become a topic that is discussed on a daily basis in laboratories all around the world. Everyone in biomedical research is now trying to incorporate this technology into some aspect of their work. And the approach has become much simplified: now researchers simply insert into a cell the Cas9 protein and specific guideRNAs for the target gene that they want to mutate and then they let the cell do the rest.
And the CRISPR-Cas9 technology has also been re-engineered in different ways to allow alternative functions to be introduced. For example, the Cas9 protein has been re-engineered so that rather than cutting DNA, the Cas9 protein now activates that region of DNA where it is directed to be the guideRNA (Click here to read more about this).
I really don’t think I’m exaggerating when I write that the potential of the technology is truly limitless (Click here to read various OPEN ACCESS articles in Cell press that highlight this fact). The only factor restricting us is our imagination.
So how can we use this technology for Parkinson’s?
While delivery of CRISPR-Cas9 technology to people with Parkinson’s is a long way off in the future (if it is possible at all), there are still numerous ways in which the technology can be applied now with beneficial effects for the community.
And researchers at the pharmaceutical company Novartis recently provided a very nice example of this by identifying positive and negative regulators of the PARKIN gene.
Here is their research report:

Title: Genome-wide CRISPR screen for PARKIN regulators reveals transcriptional repression as a determinant of mitophagy.
Authors: Potting C, Crochemore C, Moretti F, Nigsch F, Schmidt I, Manneville C, Carbone W, Knehr J, DeJesus R, Lindeman A, Maher R, Russ C, McAllister G, Reece-Hoyes JS, Hoffman GR, Roma G, Müller M, Sailer AW, Helliwell SB.
Journal: Proc Natl Acad Sci U S A. 2017 Dec 21. pii: 201711023.
PMID: 29269392 (This research report is OPEN ACCESS if you would like to read it)
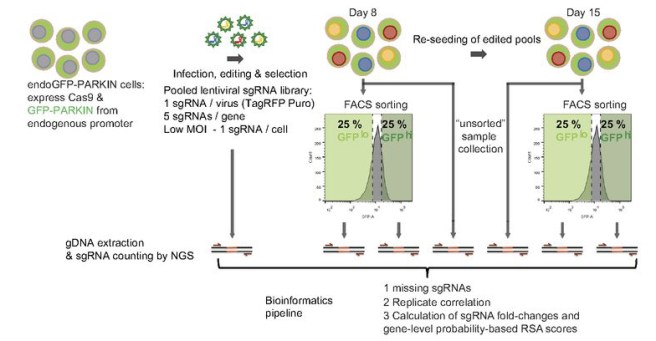
In their study, the Novartis researchers firstly engineered a cell that produced a green fluorescent protein whenever the PARKIN gene was activated. They next inserted the Cas9 protein into those cells. Then they infected the cells with a large pool of viruses – the viruses were identical except for one critical difference: each virus had a single guideRNA. The large pool of viruses collectively had enough guideRNAs to cover 18,360 genes (in fact, there were on average, five guideRNAs for each gene) – that is a lot of viruses. An ENORMOUS pool!
After infecting the cells so that each cell received just one virus, the investigators grew the cells in culture for 8 days and then sorted the cells based on their level of green fluorescent-ness. The more green the cell, the more PARKIN activation. No green fluorescence, no PARKIN activation. They also did this analysis after 15 days in cell culture to account for possible time-dependent effects of the CRISPR procedure.
Source: PNAS
The cells carrying the lowest 25% and the highest 25% of the green fluorescent-ness were analysed to determine which guideRNAs had been present in the cell. This analysis eventually gave the researchers a list of 53 positive and negative regulators of PARKIN. In the image below you can see genes that guideRNAs targets. The graphs on the left name genes that when mutated by CRISPR-Cas9 resulted in less green fluorescent protein (no PARKIN activation), and the graphs on the right name those genes that increased the levels of green fluorescent protein (more PARKIN activation). Note that PARK2 (or PARKIN) is present in both graphs on the left hand side, meaning that if PARKIN is mutated there is less activation of PARKIN.
Source: PNAS
The Novartis researchers zeroed in on three genes in particular – THAP11, HCFC1, or OGT – which when mutated, resulted in more green fluorescent protein (or more PARKIN activation). All of these three of these genes were present in both the 8 days and the 15 days analysis, suggesting that they are negative regulators of PARKIN. And given that a previous study had actually reported that depletion of THAP11 resulted in elevated levels of PARK2 (Click here for that report), the researchers focused their attention on THAP11.
The researchers began experimenting with different cell types (including human brain cell lines), and noticed that in the absence of THAP11, PARKIN levels increased, as did the levels of ubiquitin processed by PARKIN. Next they caused mitochondrial damage in the cells and found that levels of PARKIN and levels of ubiquitin processed by PARKIN increased significantly compared to control cells, indicating that a drug inhibiting THAP11 may represent a novel therapeutic approach to increasing PARKIN levels in a condition like Parkinson’s.
What does it all mean?
Parkinson’s is characterised by issues in many different biological pathways, including mitochondrial dysfunction and cellular waste disposal (autophagy). Drugs that are designed to activate genes that are involved in these processes could represent very useful tools in any future treatment of Parkinson’s.
Scientists at the pharmaceutical company Novartis has recently published research outlining the results of a massive screening study that they conducted to identify positive and negative regulators of the Parkinson’s-associated gene PARKIN, which is involved in aspects of waste disposal and maintaining healthy mitochondria. Their results provide two important pieces of information:
- Proof of principle that such screening studies can provide interesting information, and
- A list of very interesting targets for them to design inhibitors for that can be tested in models of Parkinson’s
It will be very interesting to see the follow up research to this study (hopefully in 2018, as they will have patents in place for all the interesting targets presented here, no doubt). As I mentioned above PARKIN is also involved with cancer so there is tremendous profit potential for drugs regulating this gene, and companies like Novartis will be well aware of this and pushing hard in this direction.
This research also represents a powerful application of the CRISPR-Cas9 gene editing technology for the Parkinson’s community, and I have VERY little doubt that there will be a lot more of this kind of research coming in 2018. In some cases, that research will hopefully be identifying biological pathways that could be targeted with currently available drugs that could be repurposed for Parkinson’s.
Lots of interesting therapeutic targets about to be identified. Watch this space.
The banner for today’s was sourced from the Dailypost









wow
LikeLike
I know! I couldn’t have said it any better myself 🙂
LikeLike
For me the a hope of a clinical trial can’t come to fruition fast enough.. My 36 year old daughter has Juvenile Autosomal Recessive Parkinsonism, (Parkin/Park2) diagnosed at 17 through the mapping of her DNA. Her health is not good, this disease has ravaged body. I will continue to follow this research, thanks to all the are involved in these efforts.
LikeLike