On this page we provide:
- a brief introduction to genetics
- an overview of the genetics of Parkinson’s disease
- a list of genetic regions (the “PARK genes”) associated with Parkinson’s disease
- a description of additional genetic regions that can increase risk of Parkinson’s disease
Genetics – the basics
The DNA in almost every cell of your body provides the template for making a human being. All the necessary information is encoded in that amazing molecule. The basic foundations of that blueprint are the ‘nucleotides’ – which include the familiar A, C, T & Gs – that form pairs (called ‘base pairs’) and which then join together in long strings of DNA that we call ‘chromosomes’.

The basics of genetics. Source: CompoundChem
If DNA provides the template for making a human being, it is the small variations in our individual DNA that ultimately makes each of us unique. And these variations come in different flavours: some can simply be a single mismatched base pair (also called a point-mutation or single nucleotide variant), while others are more complicated such as repeating copies of multiple base pairs.
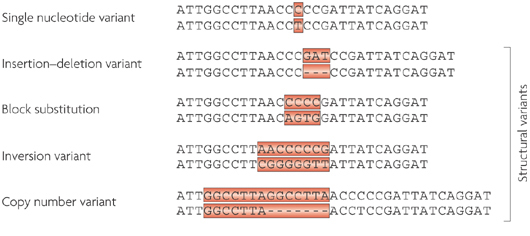
Lots of different types of genetic variations. Source: Nature
Most of the variants that we have that define who we are, we have had since conception. These are called ‘germ line’ mutations, while those that we pick up during life and that are usually specific to a particular tissue or organ in the body (such as the liver or blood), are called ‘somatic’ mutations.
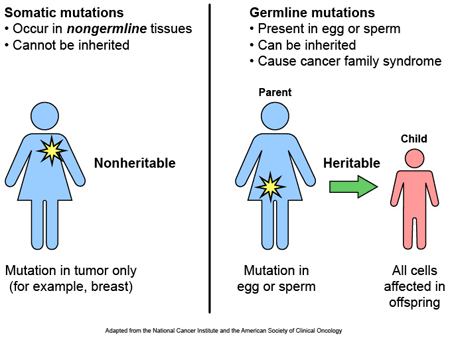
Somatic vs Germline mutations. Source: AutismScienceFoundation
In some cases, a variant has to be provided by both the parents for a condition to develop, this is called an ‘autosomal recessive‘ disease; while in other cases only need a copy of the variant to be provided by one of the parents (an ‘autosomal dominant’ disease).

Autosomal dominant vs recessive. Source: Wikipedia
Many of these tiny genetic changes infer benefits, while other variants can result in changes that are of a more serious nature.
The genetics of Parkinson’s disease
Approximately 15% of people with Parkinson disease have a family history of the condition – a grandfather, an aunt or cousin.
About 10-20% of Parkinson’s disease cases can be accounted for by genetic variations that infer a higher risk of developing the condition. In people with ‘juvenile-onset’ (diagnosed under the age 20) or ‘early-onset’ Parkinson’s disease (diagnosed under the age 40), genetic variations can account for the majority of cases, while in later onset cases (>40 years of age) the frequency of genetic variations is more variable.
For a very good review of the genetics of Parkinson’s disease – click here.
There are definitely regions of DNA in which genetic variations can increase one’s risk of developing Parkinson’s disease. These regions are referred to as PARK genes.
The PARK genes
We currently know of 23 regions of DNA that contain mutations associated with Parkinson’s disease, which have been given the name of PARK genes. The region does not always refer to a particular gene, for example in the case of our old friend alpha synuclein, there are two PARK gene regions within the stretch of DNA that encodes alpha synuclein. So please don’t think of each PARK genes as one particular gene.
In addition, there can be multiple genetic variations within a PARK gene that can infer risk of developing Parkinson’s disease. The increased risk is not always the result of one particular mutation within a PARK gene region.
A note regarding the determination of chromosomal location:
Determining chromosomal location of a PARK gene – the exact location of a PARK gene might be written as “3p22.1”. But what does this actually mean?
The “3” means chromosome 3, the “p” means p-arm (short arm) while q indicates the long arm of the chromosome. And the “22” refers to “region 2, band 2”. This is read as “two two”, not as “twenty-two”. Last “1” represents “sub-band 1”.
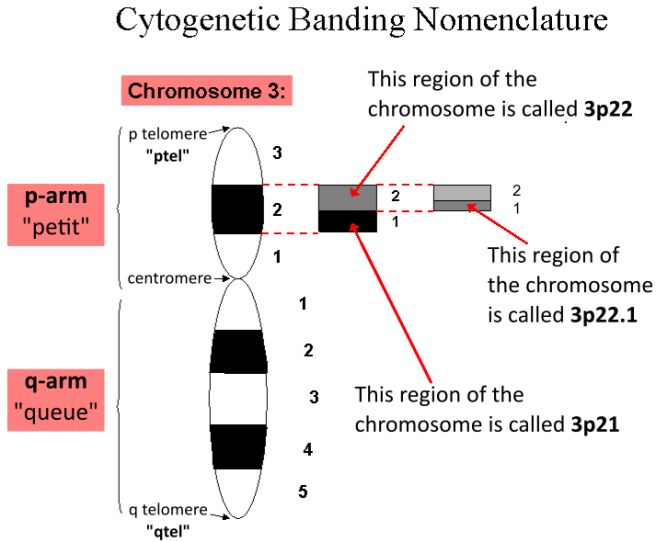
Determination of genetic location. Source: Wikipedia
The list of PARK genes:
Here is a list of genetic regions where variants within have been shown in increase the risk of developing Parkinson’s disease:
PARK1 – caused by autosomal dominant mutation (that is to say, just one copy of the variant DNA must be present) in the alpha-synuclein gene on chromosome 4q21-q22 (Gene name: SNCA; Gene ID: 6622), resulting in an early onset form of Parkinson’s disease (diagnosis before age 40).
PARK2 – caused by an autosomal recessive mutation (that is to say, two copies of the variant DNA must be present) in the Parkin gene on chromosome 6q25.2–q27 (Gene name: PRKN; Gene ID: 5071), resulting in a higher risk of juvenile onset, atypical form of Parkinson’s disease, or a classical, late-onset form of the condition.
PARK3 – caused by autosomal dominant mutation on a region of DNA on chromosome 2p13, resulting in a higher risk of classical late onset Parkinson’s disease.
PARK4 – caused by autosomal dominant mutation in the alpha-synuclein gene on chromosome 4q21–q23 (Gene name: SNCA; Gene ID: 6622), resulting in resulting in a higher risk of early-onset parkinson’s disease.
PARK5 – caused by autosomal dominant mutation in the Ubiquitin C-terminal hydrolase L1 gene on chromosome 4p13 (Gene name: UCHL1; Gene ID: 7345), resulting in a higher risk of classical late onset Parkinson’s disease.
PARK6 – caused by autosomal recessive mutation in the PTEN induced putative kinase 1 gene on chromosome 1p35–p36 (Gene name: PINK1; Gene ID: 65018), resulting in an early onset form of Parkinson’s disease (diagnosis before age 40).
PARK7 – caused by autosomal recessive mutation in the Parkinsonism associated deglycase gene on chromosome 1p36, (Gene name: DJ-1; Gene ID: 11315), resulting in an early onset form of Parkinson’s disease (diagnosis before age 40).
PARK8 – caused by autosomal dominant mutation in the leucine rich repeat kinase 2 gene on chromosome 12q12 (Gene name: LRRK2; Gene ID: 120892), resulting in a higher risk of classical late onset Parkinson’s disease.
PARK9 – caused by autosomal recessive mutation in the ATP13A2 gene on chromosome 1p36 (Gene name: ATP13A2; Gene ID: 23400), resulting in Kufor-Rakeb syndrome; atypical PD with dementia, spasticity, and supranuclear gaze palsy.
PARK10 – caused by autosomal dominant mutation in the ubiquitin specific peptidase 24 gene on chromosome 1p32 (Gene name: USP24; Gene ID: 23358), resulting in a higher risk of classical late onset Parkinson’s disease.
PARK11 – caused by autosomal dominant mutation in the GRB10 interacting GYF protein 2 gene on chromosome 2q36-27 (Gene name: GIGYF2; Gene ID: 26058), resulting in a higher risk of classical late onset Parkinson’s disease.
PARK12 – Risk factor Xq21–q25
PARK13 – susceptibility to the development of an autosomal dominant form of Parkinson’s disease may result from heterozygous mutation in the HtrA serine peptidase 2 gene on chromosome 2p13 (Gene name: HTRA2; Gene ID: 27429), resulting in a higher risk of classical Parkinson’s disease.
PARK14 – caused by autosomal recessive mutation in the PLA2G6 gene on chromosome 22q13 (Gene name: PLA2G6; Gene ID: 8398), resulting in an adult-onset dystonia-parkinsonism.
PARK15 – caused by autosomal recessive mutation in the F-box protein 7 gene on chromosome 22q12–q13 (Gene name: FBXO7; Gene ID: 25793), resulting in a higher risk of early-onset parkinsonian-pyramidal syndrome
PARK16 – caused by a mutation in a region of DNA on chromosome 1q32, resulting in a higher risk of classical Parkinson’s disease
PARK17 – caused by autosomal dominant mutation in the VPS35 gene on chromosome 16q11.2, resulting in a higher risk of classical Parkinson’s disease. Click here for an excellent overview of this PARK gene.
PARK18 – caused by autosomal dominant mutation in the EIF4G1 gene on chromosome 3q27.1, resulting in a higher risk of classical Parkinson’s disease.
PARK19 – caused by autosomal recessive mutation in the DNAJC6 gene on chromosome 1p31.3, resulting in a higher risk of juvenile onset, atypical form of Parkinson’s disease.
PARK20 – caused by autosomal recessive mutation in the SYNJ1 gene on chromosome 21q22.11, resulting in a higher risk of juvenile onset, atypical form of Parkinson’s disease.
PARK21 – caused by autosomal dominant mutation in the DNAJC13 gene on chromosome 3q22.1, resulting in a higher risk of late-onset Parkinson’s disease.
PARK22 – caused by autosomal dominant mutation in the CHCHD2 gene on chromosome 7p11.2, resulting in a higher risk of classical Parkinson’s disease.
PARK23 – caused by autosomal dominant mutation in the VPS13C gene on chromosome 15q22, resulting in an early onset form of Parkinson’s disease.
Location of your variant matters
The exact location of your variation can determine the nature of your Parkinson’s. In addition, some variants within PARK genes have been shown to increase risk of other conditions. Everything depends on where within the PARK gene a mutation actually lies.
For example, there are 10 common mutations in the PARK2 gene (Parkin) that can give rise to early-onset Parkinson’s disease. PARK2 has also been associated with different types of cancer – there are 13 cancer-related variants. Importantly, only two of the Parkinson’s related variants are associated with an increased risk of cancer (they are R24P and R275W – red+black arrow heads in the image below).

Comparing PARK2 Cancer and PD associated mutations. Source: Nature
Thus it is important to know exactly where your mutation is, if in fact you have one.
Other genetic regions that can increase the risk of Parkinson’s disease
- Glucosylceramidase beta (Gene name: GBA; Gene ID: 2629)