|
If you just ‘read the label on the can’ you would be mistaken in thinking that the International Parkinson Disease Genomics Consortium (IPDGC) is just an international group of researchers who are interested in the genetics of Parkinson’s. The group is so much more than that! I was recently invited to attend their annual meeting in Lisbon (Portugal) and it was a revelation. In today’s post, we will discuss the group, review some of the research it has produced, and I’ll explain why I am so impressed with this organisation.
|
 Source: Nothinginbiology
Source: Nothinginbiology
You will often hear researchers complaining about the state of science in the modern world. Go to any conference and they will express their concerns about the lack of funding, the lack of job security, the pressure from institutes to publish in the best journals, and the stress their jobs put on their lives (families, etc).
A career in scientific research is a privilege bestowed by society (and that should never be forgiven), but it comes with some brutal realities. It is not a 9-5 job. It is an unrelenting, all consuming beast. An abusive (failure and rejection are the norm) obsession that one cannot seem to recover from.
It’s really, really hard (I have seen marriages dissolve and lives basically ruined by it).
There is, however, a lot of discussion at research gatherings (and online) about how we can improve the ‘broken model of research’, and in today’s post I would like to share with you one group that is a shining, proactive example of such efforts.
I was invited to attend the annual meeting of the IPDGC.
And I’m really glad that I went.
What is the IPDGC?
In 2009, a worldwide collaboration was started by genetic researchers from the Unites States, United Kingdom, the Netherlands, France and Germany.
It was called the International Parkinson Disease Genomics Consortium (or IPDGC).
The initial goal was to collect genetic information – that is sequenced DNA – from enough people with Parkinson’s to allow for researchers to be confident (that is, have sufficient statistical power) to identify novel genetic risk factors for Parkinson’s.
What do you mean “novel genetics risk factors for Parkinson’s”?
To explain this, we need to understand a little something about basic genetics.
In almost every cell of your body you have DNA (or Deoxyribonucleic acid). It is a molecule that is composed of two chains which coil around each other, forming the famous double helix shape.
 Source: Pngtree
Source: Pngtree
The information stored in DNA provides the template – or the instructions – for making (and maintaining) a particular organism. All of the necessary details are encoded in that amazing molecule.
How is information stored in DNA?
In the image above, you can see strands reaching between the two chains of the DNA molecule, and it is these numerous strands that contain the information.
These strands are made up of a pair of ‘nucleotides’. These are organic molecules which contain one of four bases – the familiar A, C, T & Gs that make up our DNA code. These nucleotides pairings (called ‘base pairs’) join together in long strings of DNA.
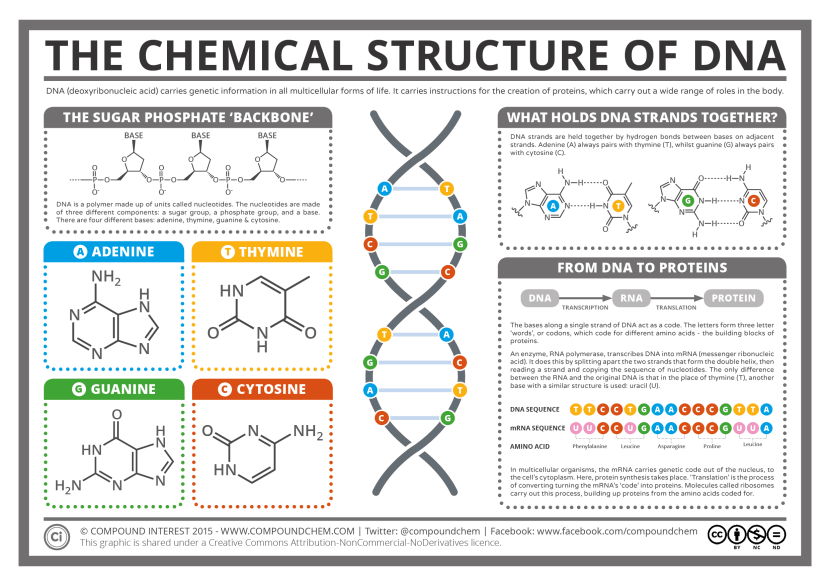 The basics of genetics. Source: CompoundChem
The basics of genetics. Source: CompoundChem
It takes a lot of instructions to make a fully functional human being, and as a result human DNA contains a lot of these base pairs – 3 billion of them. And having so many base pairs results in a lot of DNA in each cell – 2 meters of it. In order to get all that DNA crammed into the nucleus of a tiny little cell, DNA is wound up tightly into thread-like structures called chromosomes.
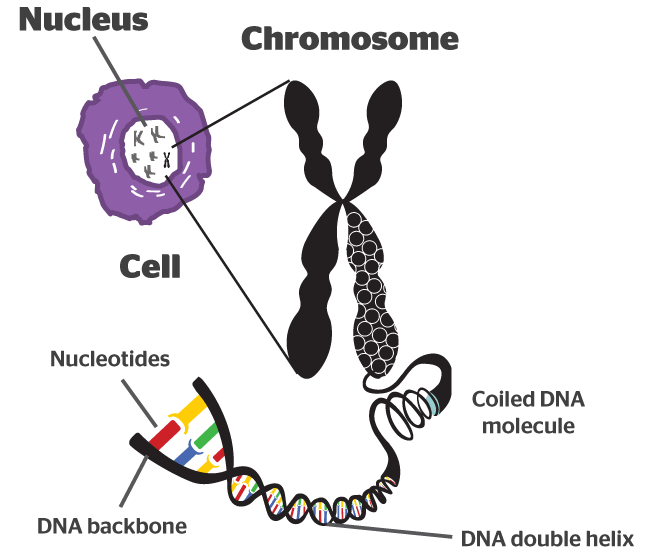 DNA in a chromosome. Source: Byjus
DNA in a chromosome. Source: Byjus
Humans have 23 pairs of chromosomes. And the genetic mutations we are talking about today can occur in any of these chromosomes.
This video does a better job of explaining DNA than I do:
Now, if DNA provides the template for making a human being, it is the small variations in our individual DNA that help to make each of us unique. Every single one of us has small genetic variations.
And these variations come in different flavours: some can simply be a single mismatched base pair (also called a point-mutation or single nucleotide polymorphism (SNP)), while others are more complicated such as repeating copies of multiple base pairs.

Lots of different types of genetic variations. Source: Nature
An example of how these variation can make us who we are is red hair. The fact that an individual has red hair results (in the majority of cases) from a variation in a region of DNA (called MC1R) on chromosome 16.
If that variation is not there: no red hair.
Most of the variants that we have that define who we are, we have had since conception. These are called ‘germ line’ mutations, while those that we pick up during life and that are usually specific to a particular tissue or organ in the body (such as the liver or blood), are called ‘somatic’ mutations.

Somatic vs Germline mutations. Source: AutismScienceFoundation
These variations can occur in regions of DNA that have no apparent use (about 90% of your DNA), but they can also appear in more functional regions (which are called genes). Genes are regions of DNA that provide the instructions for producing proteins, enzymes and pieces of regulatory RNA. These regions provide the instructions (or RNA), which can in some cases be used to make protein. Where genetic mutations fall within these regions, the resulting RNA and protein can carry a variation that will impact their function – sometimes increasing activity, sometimes preventing activity.
An important aspect of these variations in genes is that you have two copies of each gene (called alleles). One copy from your father, and one copy from your mother. Call it mother nature’s insurance policy – if one copy of a gene is faulty, you have a back up. But sometimes one copy is not enough, and this can lead to problems.
In conditions that are influenced by genetics, there are two types of inherited mutations:
- A variant has to be provided by both the parents for a condition to develop – this is called an ‘autosomal recessive‘ disease. In this case, both copies (or alleles) will be mutated, resulting in the condition having a higher chance of developing. An example of this is cystic fibrosis (a condition in which sticky mucus builds up in the lungs and digestive system).
- Only one copy of the variant needs to be provided by one of the parents for a condition to develop – this is called an ‘autosomal dominant’ disease. In this case just one allele is required for a condition to have a higher chance of developing, and a example of this is the neurodegenerative condition of Huntington’s.
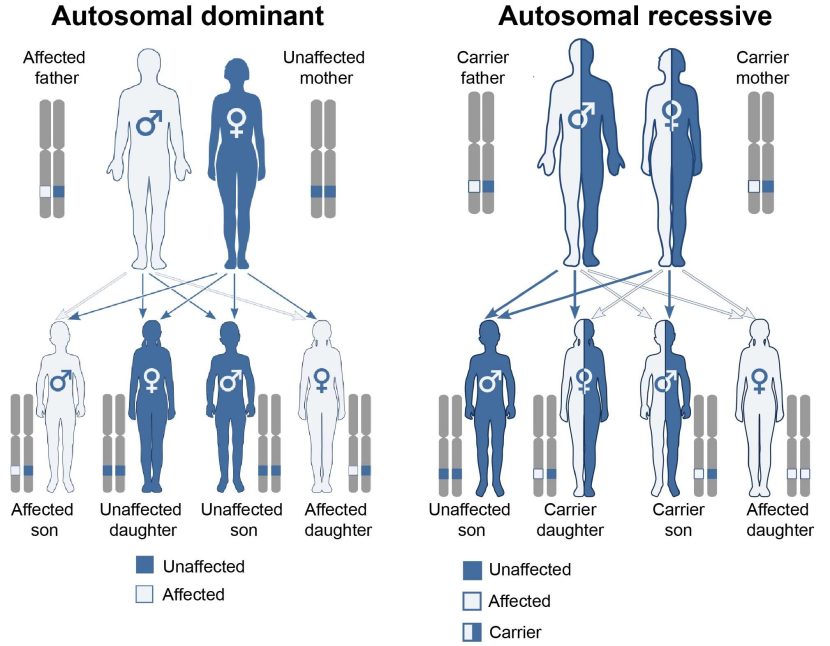
Autosomal dominant vs recessive. Source: Wikipedia
It is important to understand that most of these tiny genetic variation result in no impact on an organism. The region of DNA is not important, or biology is able to adapt and find a way around the problem.
Other variants can infer traits that may be considered beneficial, for example, some people have a genetic mutation in the CCR5 gene. HIV uses the CCR5 protein to enter into human cells. Thus, if a person has a mutation in their CCR5 gene, they are extremely unlikely to become infected by the virus.
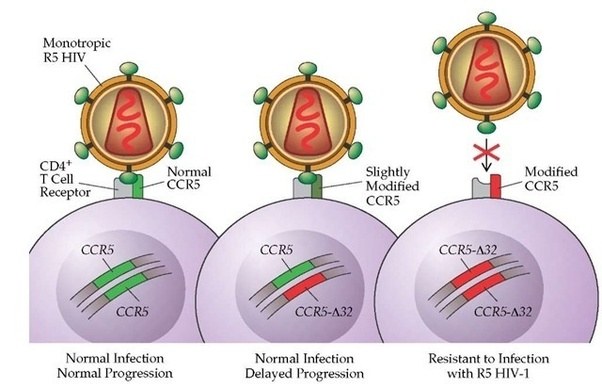 Source: Quora
Source: Quora
There are, however, some genetic variation that are of a more serious nature – leaving us potentially vulnerable to developing a condition (such as BRCA1 mutations in breast cancer). And for a long time, researchers have been interested in identifying genetic variations that are associated with Parkinson’s.
These are the genetic risk factors that we were talking about above.
So there is a genetic association with Parkinson’s?
Since it was first described by good old JP in 1817, there have been suspicions and theories of a hereditary/familial link in Parkinson’s. The condition appeared to be passed on in families – with uncles, fathers, etc also having the condition or symptoms of it.
These observations of patterns resulted in the idea that there could be a genetic link in Parkinson’s.
It was not until 1997, however, that the first genetic risk factor of Parkinson’s was discovered:

Title: Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease.
Authors: Polymeropoulos MH, Lavedan C, Leroy E, Ide SE, Dehejia A, Dutra A, Pike B, Root H, Rubenstein J, Boyer R, Stenroos ES, Chandrasekharappa S, Athanassiadou A, Papapetropoulos T, Johnson WG, Lazzarini AM, Duvoisin RC, Di Iorio G, Golbe LI, Nussbaum RL.
Journal: Science. 1997 Jun 27;276(5321):2045-7.
PMID: 9197268
In this critical study, the researchers had encountered several independent Italian and Greek families whose members exhibited a similar early onset form of Parkinson’s. The investigators found that the affected members of those families also had genetic mutations in the long arm of chromosome 4.
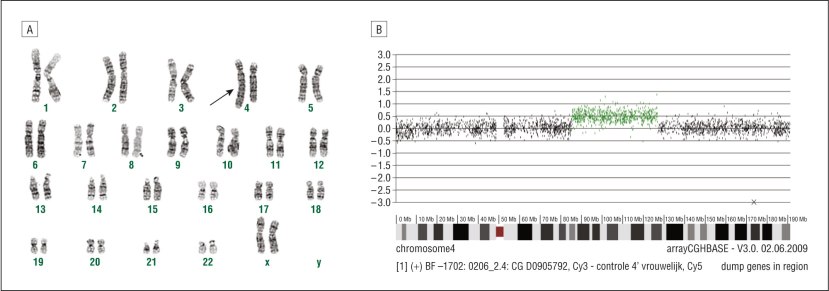
The 23 pairs of chromosomes and variants (green) in chromosome 4. Source: JAMA
It was around this same time that parts of chromosome 4 was sequenced and mapped out, and as a result the researchers were able to determine that the area covering the Parkinson’s associated genetic mutations lay in a gene called SNCA, which provides the instructions for making a protein that we here at the SoPD are rather familiar with: Alpha synuclein.
You can read more about the first Parkinson’s genetic mutations in a previous SoPD post (Click here to read that post)
Over the next few years, more genetic variations that inferred a higher risk of developing Parkinson’s were identified. These discoveries gradually resulted in a list – called the PARK genes – of regions of DNA where variations were considered risk factors for developing Parkinson’s.
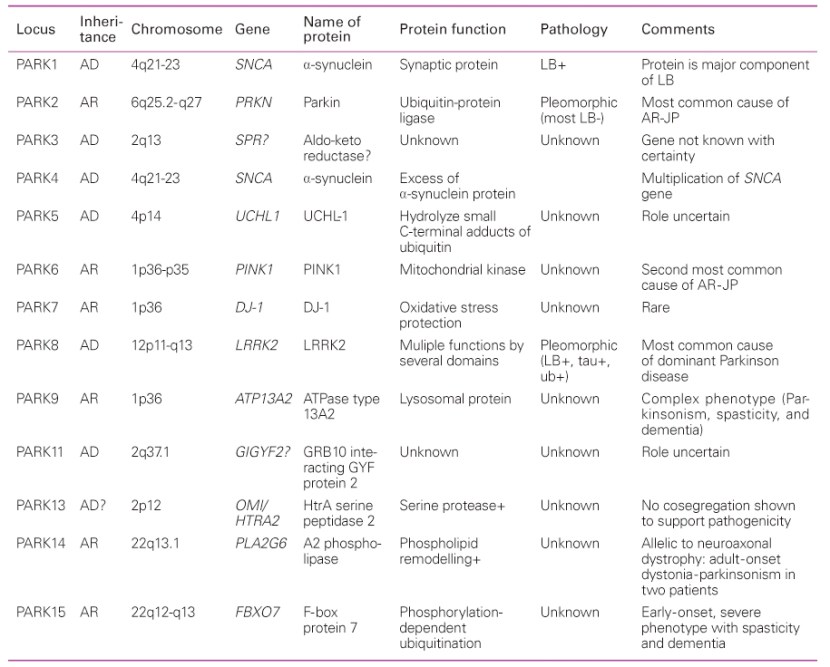
An early list of the PARK genes. Source: JKMA
All of the early PARK genes were identified by looking at families with histories of developing Parkinson’s. But using this case-by-case approach resulted in sporadic discoveries. In order to get a more thorough and complete view of the genetic component of Parkinson’s, the research community really needed BIG datasets. Gathering DNA from lots of people with Parkinson’s and sequencing it to discover all of the genetic risk factors associated with the condition.
And this is where the work of the IPDGC group really began.
Starting in 2009, they began gathering as much genetic information as they could from folks with PD.
This effort resulted in the publication of a report in 2014, which identified 28 independent risk variants for Parkinson’s:
 Title: Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease.
Title: Large-scale meta-analysis of genome-wide association data identifies six new risk loci for Parkinson’s disease.
Authors: Nalls MA, Pankratz N, Lill CM, Do CB, Hernandez DG, Saad M, DeStefano AL, Kara E, Bras J, Sharma M, Schulte C, Keller MF, Arepalli S, Letson C, Edsall C, Stefansson H, Liu X, Pliner H, Lee JH, Cheng R; International Parkinson’s Disease Genomics Consortium (IPDGC); Parkinson’s Study Group (PSG) Parkinson’s Research: The Organized GENetics Initiative (PROGENI); 23andMe; GenePD; NeuroGenetics Research Consortium (NGRC); Hussman Institute of Human Genomics (HIHG); Ashkenazi Jewish Dataset Investigator; Cohorts for Health and Aging Research in Genetic Epidemiology (CHARGE); North American Brain Expression Consortium (NABEC); United Kingdom Brain Expression Consortium (UKBEC); Greek Parkinson’s Disease Consortium; Alzheimer Genetic Analysis Group, Ikram MA, Ioannidis JP, Hadjigeorgiou GM, Bis JC, Martinez M, Perlmutter JS, Goate A, Marder K, Fiske B, Sutherland M, Xiromerisiou G, Myers RH, Clark LN, Stefansson K, Hardy JA, Heutink P, Chen H, Wood NW, Houlden H, Payami H, Brice A, Scott WK, Gasser T, Bertram L, Eriksson N, Foroud T, Singleton AB.
Journal: Nat Genet. 2014 Sep;46(9):989-93.
PMID: 25064009 (This report is OPEN ACCESS if you would like to read it)
Note the huge number of researchers involved in this collaborative effort – and there were many more names of individuals involved that were represented by various organisations mentioned in the list of authors.
In this study, the researchers analysed DNA from 13,708 people with Parkinson’s and 95,282 controls, and then validated their findings on an independent dataset of DNA from 5,353 people with Parkinson’s and 5,551 controls.
A few years later the researchers repeated this feat with DNA from even larger datasets which resulted in this study being published:
 Title: A meta-analysis of genome-wide association studies identifies 17 new Parkinson’s disease risk loci.
Title: A meta-analysis of genome-wide association studies identifies 17 new Parkinson’s disease risk loci.
Authors: Chang D, Nalls MA, Hallgrímsdóttir IB, Hunkapiller J, van der Brug M, Cai F; International Parkinson’s Disease Genomics Consortium; 23andMe Research Team, Kerchner GA, Ayalon G, Bingol B, Sheng M, Hinds D, Behrens TW, Singleton AB, Bhangale TR, Graham RR.
Journal: Nature Genet. 2017 Oct;49(10):1511-1516.
PMID: 28892059 (This report is OPEN ACCESS if you would like to read it)
In this study, the researchers conducted a GWAS comparing DNA from 6,476 people with Parkinson’s with DNA from 302,042 controls. They then compared those results with another GWAS dataset from a recent study involving 13,000 people with Parkinson’s and 95,000 controls.
Wait a minute. What is a GWAS?
A genome-wide association study (or GWAS) is an analysis of a set of genetic variants across the entire genome (or all of the DNA in your cells, including mitochondrial), and this analysis is conducted in a large pool of different individuals to see if any variants are associated with a particular trait (or medical condition). It is typically an analysis of single nucleotide polymorphisms (or SNPs; a variation in a single nucleotide). The researcher will check your DNA for the presence of a large set of single nucleotide variations, and then compare them with the results collected from other people.
 Source: Knowgenetics
Source: Knowgenetics
Now as we have discussed above, we all have these tiny mutations. But what a GWAS does, is seek to determine whether people with a particular trait (for example, constrictions in small blood vessels) have a shared single nucleotide polymorphisms, compared to people that do not have that trait. Think of the example of red hair mentioned above, a GWAS analysis of red haired people would point towards a variation on chromosome 16 (in the MC1R gene).
And the result can be presented in what is called a “Manhattan plot” – Manhattan plots get their name because they often look like the city skyline of Manhattan:
 Source: Andrewprokos
Source: Andrewprokos
Across the bottom of a Manhattan plot are the chromosomes, and each dot represents a single nucleotide polymorphisms for that particular region of the DNA. The more dots there are above that region – the more people with a variation in that region – the higher the dot in that particular location. The Manhattan plot in the image below is for an analysis of people with constrictions in small blood vessels, and what you can see is that the number of dark blue dots above chromosome 19 indicate that there are a lot of people who have this trait that also have a genetic variant in that region.
Source: Wikipedia
When researchers who conducted the Parkinson’s study from last year analysed of their data, this was the kind of Manhattan plot they observed:
 Source: PMC
Source: PMC
Immediately you can hopefully see that we are looking at a complicated set of result – there are a lot of ‘buildings’ in this Manhattan city skyline. Lots of genetic variants that could be considered risk factors for Parkinson’s.
The researchers then tested these results in a third data set of DNA from 5,851 people with Parkinson’s and 5,866 controls. From this combined analysis (totalling 26,035 PD cases and 403,190 controls), they found 17 novel risk factors for Parkinson’s (41 in total; 24 had been previously reported).
More recently, the IPDGC team has taken this analysis one step further and made this manuscript available on the preprint website BioRxiv:
 Title: Expanding Parkinson’s disease genetics: novel risk loci, genomic context, causal insights and heritable risk.
Title: Expanding Parkinson’s disease genetics: novel risk loci, genomic context, causal insights and heritable risk.
Authors: Nalls MA, Blauwendraat C, Vallerga CL, Heilbron K, Bandres-Ciga S, Chang D, Tan M, Kia DA, Noyce AJ, Xue A, Bras J, Young E, von Coelln R, Simón-Sánchez J, Schulte C, Sharma M, Krohn L, Pihlstrom L, Siitonen A, Iwaki H, Leonard H, Faghri F, Gibbs JR, Hernandez DG, Scholz SW, Botia JA, Martinez M, Corvol J-C, Lesage S, Jankovic J, Shulman LM, The 23andMe Research Team, System Genomics of Parkinson’s Disease (SGPD) Consortium, Sutherland M, Tienari P, Majamaa K, Toft M, Andreassen OA, Bangale T, Brice A, Yang J, Gan-Or Z, Gasser T, Heutink P, Shulman JM, Wood N, Hinds DA, Hardy JA, Morris HR, Gratten J, Visscher PM, Graham RR, Singleton AB for the International Parkinson’s Disease Genomics Consortium.
Journal: BioRxiv preprint
PMID: N/A (This manuscript is OPEN ACCESS if you would like to read it)
In this study, the researchers analysed DNA from 37.7K cases of Parkinson’s and 18.6K UK Biobank proxy-cases (these were samples of DNA from individuals with a first degree relative with PD, and no diagnosis or self-report of actual PD), and they compared these datasets with a control group of 1.4million (!!!) healthy individuals.
They identified 90 independent significant variants across 78 genes. The new Manhattan plot looks like this:
 As you can see, an ever more complex image of the genetics of Parkinson’s is developing. And as the number of DNA samples increases, the researchers will be able to identify very rare genetic variants that are associated with the risk of developing Parkinson’s.
As you can see, an ever more complex image of the genetics of Parkinson’s is developing. And as the number of DNA samples increases, the researchers will be able to identify very rare genetic variants that are associated with the risk of developing Parkinson’s.
So what was discussed at the latest IPDGC meeting?
The meeting was held in Lisbon (Portugal) over two days at the end of March. About 100 researchers attended, and representatives from various government and funding bodies (such as Michael J Fox Foundation & the Parkinson’s Foundation).
 Dr Jim Beck from the Parkinson’s Foundation presenting their new PD GENEration initiative
Dr Jim Beck from the Parkinson’s Foundation presenting their new PD GENEration initiative
The morning of the first day focused on ongoing efforts around the world to collect DNA and clinical datasets. And when I say around the world, I mean around the world! DNA is being collected in Africa, India, South America, and Asia, and reports of the work were discussed by the group. Very interesting differences between various geographical regions are starting to appear (the topic of a future SoPD post). The current goal of the IPDGC group is to get 100K cases of Parkinson’s.
During the afternoon session, the meeting shifted to ongoing clinical-genetic initiatives (of which there were quite a few). These projects were attempting to find associations between genetic variations and clinical outcomes. They included efforts to identify the genetics underlying atypical Parkinsonisms (such as Mulitple Systems Atrophy and Progressive Supranuclear Palsy) and also REM sleep behaviour disorder (which is considered an early risk factor for developing Parkinson’s).
 The 2019 IPDGC meeting. Source: PredictPD
The 2019 IPDGC meeting. Source: PredictPD
Of particular interest, however, was the genetic analysis of the progression of Parkinson’s.
With the huge datasets that are being generatated by these and other researchers, an obvious question to ask is whether there are genetic variations that can infuence the speed of Parkinson’s progression.
And the answer appears to be “Yes, there are”, and the IPDGC team have provided this manuscript on BioRxiv highlighting some examples of these genetic variants that influence Parkinson’s progression:
 Title: Genome-wide association study of Parkinson’s disease progression biomarkers in 12 longitudinal patients’ cohorts
Title: Genome-wide association study of Parkinson’s disease progression biomarkers in 12 longitudinal patients’ cohorts
Authors: Iwaki H, Blauwendraat C, Leonard HL, Kim JJ, Liu G, Maple-Grødem J, Corvol J-C, Pihlstrøm L, van Nimwegen M, Hutten SJ, Khanh-Dung HN, Rick J, Eberly S, Faghri F, Auinger P, Scott KM, Wijeyekoon R, Van Deerlin VM, Hernandez DG, Gibbs JR, Chitrala KN, Day-Williams AG, Brice A, Alves G, Noyce AJ, Tysnes O-B, Evans JR, Breen DP, Estrada K, Wegel CE, Danjou F, Simon DK, Andreassen O, Ravina B, Toft M, Heutink P, Bloem BR, Weintraub D, Barker RA, Williams-Gray CH, van de Warrenburg BP, Van Hilten JJ, Scherzer CR, Singleton AB, Nalls MA
Journal: BioRxiv preprint
PMID: N/A (This manuscript is OPEN ACCESS if you would like to read it)
In this study, the researchers used individual data from not one, but 12 large longitudinal cohort studies. The data involved both genetic and clinical information on 4,093 people with Parkinson’s (the data included over 25,000 clinical assessments). Using this large database, the investigators identified nine associations between known Parkinson’s risk variants and more severe motor or cognitive symptoms.
But more importantly, they also found a genetic variant (Rs382940(T>A)) within the SLC44A1 gene which was associated with faster progress (reaching Hoehn and Yahr stage 3 or higher faster). It is interesting to note that the SLC44A1 gene has not previously been associated with increased risk of developing Parkinson’s. This variant is only associated with the speed of progression. Such information may be very useful in clinical trials for disease modifying therapies, as individuals with this genetic variant will need to be identified (and potentially excluded) so as to not to potentially bias the results.
The researchers also found a genetic variant (Rs61863020(G>A)) in the ADRA2A gene which was associated with a lower prevalence of insomnia at the baseline assessment. This kind of genetic research is demonstrating that tiny variations in our DNA are not only associated with our risk of developing Parkinson’s, but also associated with the nature of the Parkinson’s.
Have they found any genetic variations that reduce the risk of Parkinson’s?
No. Not yet.
They are hoping, however, that some kind of protective variants will begin to appear in the data once they have a large enough dataset (that goal of 100K should be enough). Such information could be hugely informative not only from the stand point of underlying biology of PD, but also with regards to future therapeutics.
And what happened on day two of the meeting?
The second day focused more on how genetics is influencing the biology of Parkinson’s-associated protein, such as alpha synuclein and LRRK2.
There were numerous presentations on efforts to identify genetic ‘modifiers’ of these proteins. Some of the research that was presented during these sessions will be part of future SoPD posts, so to save space I’m going to skip them here.
During the afternoon, however, there was session wrapping up the meeting. And it was in this part of the event that Dr Margaret Sutherland (Program Director for Neurodegeneration at the National Institute of Neurological Disorders and Stroke (NINDS) – a part of the U.S. National Institutes of Health (NIH)) spoke. She provided an overview of the Accelerating Medicines Partnership (AMP) program being developed by the NIH for Parkinson’s (AMP PD).
 Source: NIH
Source: NIH
Launched in 2014, the initial projects of the AMP project have focused on Alzheimer’s disease, type 2 diabetes, and the autoimmune disorders rheumatoid arthritis and lupus. And now it is Parkinson’s turn.
AMP PD will focus on identifying and validating promising biomarkers that may be useful in tracking the progression of Parkinson’s and could serve as biological targets for the development of new drugs. Critically, the initiative will bring together various partners – including Celgene, GlaxoSmithKline, The Michael J. Fox Foundation for Parkinson’s Research, Pfizer, Sanofi, and Verily – and pool together huge amounts of data from many different sources, from past clinical trials and the Parkinson’s Progression Markers Initiative (PPMI) to vast genetics databases.
A beta version of the AMP PD website (‘knowledge portal’) is currently being beta tested with the final version hopefully rolling out later this year (Click here to read more about this ambitious program).
It sounds like the AMP PD knowledge portal will be an amazing resource.
Sounds like a fantastic meeting. Is all of this data and information the reason why you were so impressed with this meeting/organisation?
No.
While the amount of work the group is producing is hugely impressive, it was minor compared to the culture change that they are developing. The level of collaboration and data sharing within this consortium is unbelievable. Everyone knows who is working on what topics and they all seek to help each other with tools and ideas. All of the manuscripts generated by the group are firstly shared with everyone within the consortium to get feedback and thoughts, before it is submitted for publication.
And before the manuscripts are published in a scientific journal, they are also made available to the wider research community via the preprint website BioRxiv. Note that several of the studies discussed in this post are available to the public on BioRxiv and still awaiting formal publication.
A regular message at the meeting was “data sharing for the common good”.
And I can’t tell you how refreshing and inspiring it was to hear this. So much of the modern research community are burdened with the paranoia about ‘being scooped’ (beaten to publication by competitors) that they are too terrified to share anything (manuscripts, data, tools, or even ideas). To be fair, their careers are on the line, with their research institutes judging them on how well they publish and how much research funding they can attract.
But the culture being encouraged by the IPDGC group should be evaluated and – where possible – replicated.
And no where was this collaborative energy more evident than the final wrapping up session.
During each section of the meeting, action points were being written up on a white board, and in the wrapping up part of the meeting, Andy Singleton – who was chairing the session – was asking for volunteers to work on each of the action points. The atmosphere in the room was electric. Individuals were volunteering to take on tasks, and there was a dynamic discussion of ideas.
‘Inspiring’ – is the best word to describe it.
And all for a ‘common good’.
Interesting. So summing up – what does it all mean?
No wait, I’m not done yet.
There’s one other aspect of the meeting I wanted to hightlight: The consortium also encourages the movement of students between labs, to learn new skills and share ideas. And this may not sound like much of an idea until you realise that some of the students are going to Nigeria to be part of the African Parkinson’s genetics efforts, and students from Nigeria are going to the US (or Europe) to learn new skills and best practises that they can then take back. Similarly, students are encouraged to go to South America and other parts of the world, and individuals from those regions travel as well.
I was REALLY impressed by this kind of collaborative atmosphere. It gave new meaning to the idea of ‘common good’.
So what does it all mean?
Few members of the affected Parkinson’s community have probably even heard of the IPDGC and their efforts.
And perhaps members of the Parkinson’s research community take for granted a lot of the data being generated by the consortium. But I think a great deal could be learnt from this group – and I’m not referring to genetics.
Before attending the annual meeting of the International Parkinson Disease Genomics Consortium (IPDGC), I was aware of the huge amount of data being generated by the group – they have been a proactive member of most of the research focused on the genetics of Parkinson’s to date. But what I discovered at the IPDGC meeting was something else entirely and I found terribly inspiring. A collaborative collection of intelligent individuals proactively working together to provide the Parkinson’s world with an amazing asset.
I am extremely grateful to have had the opportunity to attend this meeting, and I wanted to share my observations and experience with readers.
The attendees of the 2019 International Parkinson Disease Genomics Consortium (IPDGC) meeting:
 (I didn’t notice at the meeting, but Prof Steve Finkbeiner (second from the left, front row) is wearing an All Blacks jersey!)
(I didn’t notice at the meeting, but Prof Steve Finkbeiner (second from the left, front row) is wearing an All Blacks jersey!)
EDITOR’S NOTE: The author of this blog has no direct association with the IPDGC group. He was invited to attend the two day meeting before the ADPD 2019 meeting (immediately following the IPDGC event). The meeting was supported by the Parkinson’s Foundation. The IPDGC group has not requested the production of this post, nor were they made aware of it before publication. It was generated simply because the author was so impressed with the meeting and with the culture being encouraged by the overall group, that he thought readers may be interested.
The banner for today’s post was sourced from IPDGC





Awesome collaboration! Nice looking group. The people in the back row look kinda shy except for one really tall guy in the middle …
LikeLike
Yes, he even stands out in the crowd
LikeLike
Simon…thank you so much for writing this, it was wonderful that you were able to attend the meeting, I have to say I was incredibly touched by your article – it has done the rounds through our IPDGC mailing list, and I think we all feel a little proud. Hope you’ll make it again next time!
LikeLike