
The genetics of any disease is very complicated. We are, however, gradually identifying the genetic mutations/variations that are associated with Parkinson’s disease and coming to understand that role of those genes in the condition. This week, researchers have identified a mutation underlying one form of Parkinson’s disease, which is associated with the name PARK16.
In this post we will review what the scientists have found and what it means.

A map of some of the genetic interactions associated with Parkinson’s disease. Source: Pubmed
As the image above demonstrates the genetic interactions underlying some forms of Parkinson’s disease are extremely complicated. And it is important to note, dear reader, that that schematic provides only a partially completed picture. It maps out only a portion of the interactions that we know of, and we can only guess at the interactions that we don’t know of. Complicated right?
Approximately 10-15% of cases of Parkinson’s disease are associated with a genetic variation in the DNA that renders an individual vulnerable to the condition.
The region of DNA in which a mutations occurs is called the ‘Locus’. There are more than 20 loci (these regions of mutations) now associated with Parkinson’s disease. The loci are referred to as ‘PARK genes’.
What are the PARK genes?
Below is a table of the first 15 PARK genes to be associated with Parkinson’s disease:
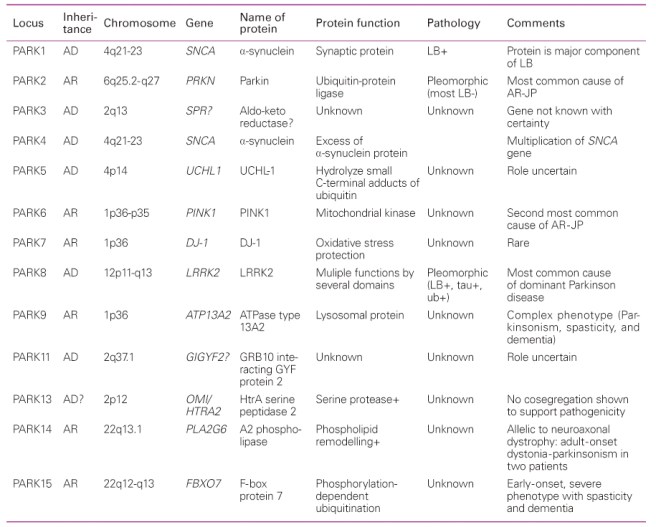
A list of the PARK genes. Source: JKMA
The PARK genes in the table are numbered 1 to 15 (16-20 are not mentioned here), and their genetic location is indicated under the label ‘Chromosome’ (this tells us which chromosome the locus is located on and where on that chromosome it is). The specific gene and protein that are affected by the mutation are also labelled (for example the gene (and protein) associated with PARK8 is Lrrk2). It is interesting to note that the gene responsible for making the protein alpha synuclein (SNCA) has two PARK gene loci within it (PARK 1 and PARK4), further emphasizing the importance of this gene in the disease.
You may also notice that there are a lot of unknowns under the labels ‘protein function’ and ‘Pathology’ (with regards to Parkinson’s disease), this is because we are still researching these genes. Furthermore, PARK3 and PARK11 both have question marks beside the genes associated with these loci, indicating that we are still not sure if these are the genes responsible for the dysfunction we observed in these forms of Parkinson’s disease.
Obviously the PARK genes list is a work in progress.
That said, this week researchers from the University of Tehran (Iran) published a report about the gene they believe is responsible for the dysfunction associated with PARK16 mutations:

Title: Mutation in ADORA1 identified as likely cause of early-onset parkinsonism and cognitive dysfunction.
Author: Jaberi E, Rohani M, Shahidi GA, Nafissi S, Arefian E, Soleimani M, Moghadam A, Arzenani MK, Keramatian F, Klotzle B, Fan JB, Turk C, Steemers F, Elahi E.
Journal: Mov Disord. 2016 May 2.
PMID: 27134041
The researchers had two siblings (brothers) referred to them that had been diagnosed with early onset Parkinson’s disease (2 siblings from a family of 8 children). Both of the siblings were in their early 30s, but had exhibited Parkinson’s-like features since their early 20s. They had responded to L-dopa therapy, but involuntary movements (L-dopa-induced dyskinesias) had started to appear after just 2 years of treatment.
Naturally the researchers were keen to determine if there was a genetic reason for this situation. To this end, they conducted whole genome analysis to determine what genetic variations the two siblings shared.
They took DNA from white blood cells of the 10 family members (two parents and eight children), and sequenced the genomes for analysis. What they found was two regions of DNA that were the same in the two affected siblings, but different in the rest of the family. In one of these regions was in the gene ADORA1, which encodes a receptor for a particular protein that can influence dopamine release. Importantly, the ADORA1 gene is located within the domain of the PARK16 locus.
When the researcher checked the sibling’s genetic variation inside the ADORA1 gene on a database of 60,000 normal individuals, they found only one other individual who was partially affected by it, suggesting that this mutation is very rare. Based on these findings, the researchers concluded that variations in ADORA1 may explain some of the cases of PARK16 -associated Parkinson’s disease.
So what does it all mean?
It means that we have another piece of the puzzle, and each week other pieces are falling into place. ADORA1 may not be the only genetic variant within the PARK16 locus, but it will explain some cases of PARK16 Parkinson’s disease. Next we need to work out what the variation does to the gene function of ADORA1.
And that will hopefully be a future blog post.
4 thoughts on “Finding PARK16”