
|
This week multiple research groups at the University of Oxford and Boston-based FORMA Therapeutics announced a collaboration to identify, validate and develop deubiquitinating enzyme (DUB) inhibitors for the treatment of neurodegenerative conditions, like Parkinson’s. But what exactly are DUB inhibitors? And how do they work? In today’s post, we will answer these questions, look at what the new collaboration involves, and look at what else is happening with DUB inhibitors for Parkinson’s. |

Source: Blog4dubstep
Dubstep is a genre of electronic dance music that originated in South London in the late 1990s. Only recently -in the 2010s – has the culture really become more mainstream. And while I have a hard time appreciating the heavy bass music (man, I am becoming a grumpy old man before my time), it is amazing to watch some of the dancers who robotically embody this form of music:
The guy on the right is named Marquese Scott. Sometimes he simply defies the laws of physics.
The title of today’s post is a play on words, because rather than doing ‘Dubstep’ we are going to be discussing how to ‘DUB-stop’.
Researchers in Oxford have recently signed an agreement with a US company to focus resources and attention on a new approach for tackling neurodegenerative conditions, including Parkinson’s.
What they are proposing is a complicated biological dance.
Their idea: to stop deubiquitinating (DUB) enzymes.
What are deubiquitinating enzymes?
Deubiquitinating enzymes are a very large group of proteins (there are 102 in humans) that function as enzymes (surprise!).
They are critically involved in a process that is called ubiquitination.
What is ubiquitination?
Ubiquitination is the process of adding ubiquitin to a protein.
This process can affect proteins in many different ways. Ubiquitin can:
- Label them for removal/disposal
- Alter their location within a cell
- Change their level of activity
- Promote/prevent interactions with other proteins
Ubiquitin is a marvellous little protein that – as the name suggests – is ‘ubiquitous’ in all cells, and as suggested above it affects all aspects of cell biology.
The structure of ubiquitin. Source: Wikipedia
The most studied process associated with ubiquitination is the marking of particular proteins for disposal.
There are primarily two ways that cells dispose/recycle damaged or old stuff. And both of them involve ubiquitination.
The first is called the Ubiquitin Proteasome Pathway.
Source: 2bscientific
A protein labelled with a lot of ubiquitin will be identified and sent off for disposal and be broken down in a structure called the proteasome. For a more thorough explanation of the Ubiquitin Proteasome pathway, please watch this video kindly provided by the Scottish Enterprise:
A second process by which cells dispose of rubbish is called autophagy.
Autophagy (from the Ancient Greek αὐτόφαγος autóphagos, meaning “self-devouring”) is an absolutely essential function in a cell. Without autophagy, old proteins would pile up making the cell sick and eventually causing it to die. Through the process of autophagy, the cell can break down the old protein, clearing the way for fresh new proteins to do their job.

The process of autophagy. Source: Wormbook
Waste material inside a cell is collected in membranes that form sacs (called vesicles). These vesicles then bind to another sac (called a lysosome) which contains enzymes that will breakdown and degrade the waste material – the same way enzymes in your washing powder break down muck on your dirty clothes). The degraded waste material can then be recycled or disposed of by spitting it out of the cell.
There are different types of autophagy, and one is thought to be rather critical in the case of Parkinson’s.
It is called mitophagy and it involves the disposal of old or dysfunctional mitochondria.
Mitochondria – you may recall from previous SoPD posts – are the power stations of each cell. They help to keep the lights on. Without them, the party is over and the cell dies.

Mitochondria and their location in the cell. Source: NCBI
You may remember from high school biology class that mitochondria are tiny bean-shaped objects within the cell. They convert nutrients from food into Adenosine Triphosphate (or ATP). ATP is the fuel which cells run on. Given their critical role in energy supply, mitochondria are plentiful (some cells have thousands) and highly organised within the cell, being moved around to wherever they are needed.
Like you, me and all other things in life, mitochondria have a use-by date.
As mitochondria get old and worn out (or damaged) with time, the cell will dispose of them via mitophagy.
There are two Parkinson’s-associated proteins that play important roles in mitophagy. They are named PARKIN and PINK1.
PINK1 acts like a kind of handle on the surface of mitochondria. In normal, healthy mitochondria, the PINK1 protein attaches to the surface of mitochondria and it is slowly absorbed until it completely disappears from the surface and is degraded. In unhealthy mitochondria, however, this process is inhibited and PINK1 starts to accumulate on the outer surface of the mitochondria.
If PINK1 a handle on the surface of the mitochondria, then PARKIN is the flag that likes to hold onto the PINK1 handle. While exposed on the surface of mitochondria PINK1 starts grabbing the PARKIN protein. This pairing activates PARKIN which starts attaching ubiquitin to molecules on the surface of the mitochondria. When enough ubiquitin is present, the cell is alert to the fact that this particular mitochondrion (singular) is not healthy and needs to be removed.

Pink1 and Parkin in normal (right) and unhealthy (left) situations. Source: Hindawi
Think of both autophagy and the proteosome pathway as the waste disposal/recycling mechanisms of the cell.
And understand that ubiquitin is a critical to both processes.
How are deubiquitinating enzymes involved in these process?
The biological world is all about balance.
For every enzyme, like PARKIN that likes to attach ubiquitin to something, there needs to be an enzyme that does the opposite (removes ubiquitin from stuff).
And this is – as the label on the can suggests – what ‘deubiquitinating’ enzymes do. They are actively involved with the removal of ubiquitin from proteins.
Thus, while there are some enzymes rapidly attaching ubiquitin to proteins, there are equally enzymes removing it. This gives us balance. Without deubiquitinating enzymes, ubiquitin attaching enzymes would simply run wild. And without ubiquitin attaching enzymes, deubiquitinating enzymes would prevent any ubiquitination from occurring resulting in all activity in the cell coming to a complete halt.
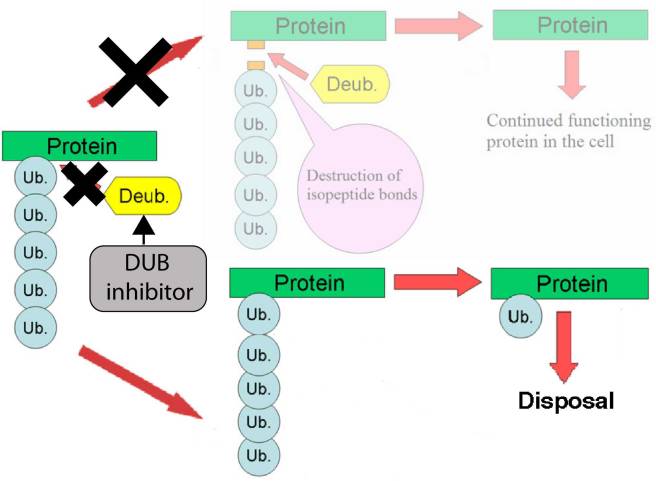
This balance also means that the difference between a protein inside the cell being allowed to continue functioning or being sent off for disposal depends on how much ubiquitin is attached to it.
As illustrated in the image below deubiquitinating enzymes (Deub; labelled in yellow) remove ubiquitin (Ub) from proteins. But if they don’t take enough off, the protein is sent off for disposal.
Source: Wikipedia
Ok, but why would researchers be interested in deubiquitinating enzymes?
A while back, some clever researchers realised that by blocking or enhancing specific deubiquitinating enzymes, one could potentially tip the balance in favour of either reducing or increasing levels of a particular protein.
For example, if there is a particular deubiquitinating enzyme that removes ubiquitin from a protein like Parkinson’s-associated Alpha Synuclein, one could design a drug (a DUB inhibitor) targeting that deubiquitinating enzyme which would block it from removing ubiquitin from alpha synuclein. This would result in more ubiquitin remaining on alpha synuclein protein, thus increasing the chances of its removal and disposal.
Deubiquitinating enzyme (DUB) inhibitors represent a method of increasing the removal of a particular protein from a cell.
For a good review on the topic of Deubiquitinating enzyme – Click here.
Interesting idea. Are there any associations between deubiquitinating enzymes and Parkinson’s?
Yes, there are.
In fact, genetic variations in two particular DUB enzymes genes can lead to the development of Parkinson’s.
Those two genes are:
- Ubiquitin C-terminal hydrolase L1 (also known as PARK5) – genetic variations in this region of DNA result in a higher risk of classical late-onset Parkinson’s. UCHL1 is probably the most studied DUB, because it has associations with both neurodegenerative conditions and the progression of malignancies. UCHL1 is extremely abundant in all neurons (remarkably it alone accounts for 1-2% of total brain protein), and it is also found in Lewy bodies (Source). Curiously, one genetic mutation in this gene is associated with increased risk of Parkinson’s (Source), while another genetic variant in this gene is actually associated with a reduced risk of developing Parkinson’s (Source).
- Ubiquitin specific peptidase 24 (also known as PARK10) – genetic variations in this region of DNA also result in a higher risk of classical late onset Parkinson’s disease. This enzyme is known to remove ubiquitin from damage-specific DNA-binding protein 2 (DDB2). By removing the ubiquitin from DDB2, the enzyme increases the stability of DDB2. DDB2 is involved in DNA damage recognition, which means that an unstable version of DDB2 could result in increased risk of high levels of damaged DNA.
It is remarkable to note that of the 23 PARK genes (genes associated with Parkinson’s), two of them are DUBs.
Is there any research of DUB inhibitors in Parkinson’s?
Nothing in clinical trials (yet), but there has recently been some very interesting pre-clinical research:
Title: The mitochondrial deubiquitinase USP30 opposes parkin-mediated mitophagy.
Authors: Bingol B, Tea JS, Phu L, Reichelt M, Bakalarski CE, Song Q, Foreman O, Kirkpatrick DS, Sheng M.
Journal: Nature. 2014 Jun 19;510(7505):370-5.
PMID: 24896179
In this study, the researchers found that by artificially increasing levels of the deubiquitinating enzyme ‘USP30’ in cells, the ubiquitin attached by PARKIN onto damaged mitochondria was rapidly removed. This effectively blocked the ability of PARKIN to cause mitophagy to occur. When the investigators conducted the reverse experiment – reducing USP30 activity – they found enhanced levels of mitochondrial degradation in neurons.
Next, the researchers reduced USP30 levels in flies treated with a neurotoxin (paraquat). They found that lowering USP30 levels reduced dopamine cell loss, rescued motor function, and improved overall survival (compared to neurotoxin-only treated flies). These results suggested to the researchers that USP30 inhibition could be potentially beneficial for Parkinson’s.
These results have been replicated by independent research groups (Click here and here to read that research), and supported by other research looking at USP30 and PARKIN (Click here and here to read more about this). And there are now efforts to design drugs based on this interaction.
A biotech company here in Cambridge (UK) called Mission Therapeutics are now pre-clinically testing USP30 inhibitors for Parkinson’s (Click here to read more about this). Those efforts are supported by the Michael J Fox Foundation (Click here to read the press release).
The company presented some results at the Society for Neuroscience meeting in November 2017, which highlighted a USP30 inhibitor compound called MTX090795 that provided a 40% rescue of dopamine neurons (compared to placebo treatment) when given to mice that were treated with a neurotoxin (MPTP) (Click here to read the abstract).
The company is now conducting further pre-clinical investigations of MTX090795 for Parkinson’s. There is currently no indications as to when (or if) they will take this drug to clinical trials.
But USP30 inhibition is not the only DUB that has generated interest in the Parkinson’s research community:
Title: Deubiquitinase Usp8 regulates α-synuclein clearance and modifies its toxicity in Lewy body disease.
Authors: Alexopoulou Z, Lang J, Perrett RM, Elschami M, Hurry ME, Kim HT, Mazaraki D, Szabo A, Kessler BM, Goldberg AL, Ansorge O, Fulga TA, Tofaris GK.
Journal: Proc Natl Acad Sci U S A. 2016 Aug 9;113(32):E4688-97.
PMID: 27444016 (This report is OPEN ACCESS if you would like to read it)
In this study, the researchers reported that the deubiquitinating enzyme USP8 is possibly an interesting target for therapeutic invention in Parkinson’s. The investigators firstly found that USP8 protein is present in Lewy bodies – the circular clusters of alpha synuclein that characterise the Parkinsonian brain.
USP8 (red) & alpha synuclein (green) in a Lewy body. Source: NCBI
The investigators next looked at what USP8 is doing in normal healthy cells, and they found that it is very actively removing ubiquitin from alpha synuclein protein. When they increased levels of USP8 (that is, increased the removal of ubiquitin from alpha synuclein), they found that the rate of alpha synuclein disposal was reduced. And when they conducted the reverse experiment – reducing levels of USP8 – they found that the rate of alpha synuclein disposal was increased.
With this result, the researchers next wanted to determine if they could reduce levels of USP8 and rescue a model of Parkinson’s. They chose a genetically engineered fly model (which has been created to produce high levels of human mutant alpha synuclein (A53T)). These flies have problems with their eyes due to the high levels of mutant alpha synuclein protein, but when the researchers reduced the levels of USP8 in these flies, the problems with the eyes were completely rescued. These results left the investigators concluding that inhibiting USP8 could be an interesting future therapeutic avenue for Parkinson’s.
And yes, there is a company developing inhibitors for USP8.
It is called Hybrigenics Pharma.
This Paris-based biotech company is primarily focused on different kinds of cancer, and their USP8 research is still in the pre-clinical stage. But if they discover a viable inhibitor of USP8 it would be very interesting to test it in models of Parkinson’s.
So what does the new collaboration in Oxford involve?
Under the terms of the agreement, FORMA Therapeutics and 5 departments (including the Oxford Parkinson’s Disease Centre) at the University of Oxford have signed a multi-year research collaboration to advance the development of treatments for neurodegenerative diseases (Click here for the press release).
Specifically, FORMA will “fund a multi-year research program at the University of Oxford focusing on DUBs implicated in the pathogenesis of neurodegenerative” conditions. And in exchange for this, FORMA will be granted the right to develop and commercialise the DUB inhibitors studied under the agreement.
The collaboration creates a strong synergy between FORMA Therapeutics’ expertise in small molecule drug design and development with the University of Oxford’s focus on neurodegenerative conditions and neuronal cell biology.
It will be very interesting to see what they find.
So what does it all mean?
Deubiquitinating (DUB) enzymes prevent proteins from being sent off for disposal/recycling.
By inhibiting DUBs, researchers are hoping to increase the rate at which specific disease-associated protein get removed from cells, thus improving the health of the cell. A new collaboration has been formed between a US-based biotech firm and research groups at Oxford University to identify and develop DUB inhibitors for neurodegenerative conditions, like Parkinson’s.
While this is a very interesting and exciting approach for developing novel therapeutics, many of these DUBs may have additional roles in our bodies other than just regulating particular proteins associated with neurodegenerative conditions. For example, USP8 is apparently important in the trafficking of both normal and misfolded alpha synuclein, which could result in off-target side effects. These additional roles will need to be explored – and the DUB inhibitors being developed at the moment may be useful tools for that research.
If very specific inhibitors of USP30 can be designed and developed, however, this DUB inhibitor approach could be very useful for people with particular genetic mutations, such as PARKIN variants. Given that these individuals have reduced activity of PARKIN, the balance between DUB enzymes and PARKIN is off-kilter. By lowering the levels of USP30, the balance of the PARKIN-USP30 dance could be brought back into alignment. This would not require a complete inhibition of USP30 – just enough to have an effect.
A difficult dance.
Let’s see how well the researchers do.
The banner for today’s post was sourced from Pinterest









Great video. Interesting post
LikeLike
Thanks Dkdc – glad you liked it. And the dubstep dancers are very clever: https://www.youtube.com/watch?v=NzArlTkwCN8
LikeLike
could you also tell us about ferroptosis please?
LikeLike
Hi Daniel,
Thanks for the interesting question. I have actually been thinking about doing another post on iron processing as a follow up to a previous post on the topic, which looked at a couple of iron chelators (https://scienceofparkinsons.com/2017/04/19/iron-life-force-and-parkinsons-disease/). There were a couple of updates to that post (at the bottom of that page), but the topic certainly deserves another post.
Ferroptosis is an iron-dependent programmed cell death pathway. It starts with the depletion of glutathione – which is one of the major antioxidants of neurons – in cells. This is followed up by a lot of lipid peroxidation, which results in oxidative stress, lots of free radicals, and ultimately cell death. Early preclinical research suggests that specific ferroptosis inhibitors (such as ferrostatin-1) are beneficial in models of Parkinson’s (https://www.ncbi.nlm.nih.gov/pubmed/27189756 & https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5327502/). I am not aware of an specific inhibitors being tested in clinic (happy to be corrected on that), but it is still early days for those inhibitors. Curiously, the chemical dopamine – the neurotransmitter which is so severely depleted in Parkinson’s – has been shown to blocked ferroptosis (https://www.ncbi.nlm.nih.gov/pubmed/27793671).
It is a really interesting topic. Probably worth writing a post about in the near future. I will add it to the list – thanks!
Thanks,
Simon
LikeLike