|
There is a great deal of interest in genetic risk factors in Parkinson’s at the moment. A number of companies are providing direct-to-consumer services which provide individuals with some information about their family history and whether they have any of the more common genetic variations that are associated with medical conditions, like Parkinson’s. Recently a new genetic data company has started – called Nebula Genomics – and they are offering a slightly different kind of service. While many of the direct-to-consumer genetic companies have a business model that involves selling on genetic information to third parties, Nebula is offering a more patient-empowering option. In today’s post, we will discuss the genetics of Parkinson’s, what Nebula Genomics is offering, and how this new service could be useful for the Parkinson’s community.
|
 Prof George Church. Source: Biospace
Prof George Church. Source: Biospace
Professor George Church is a person most readers will have never heard of.
He is the Robert Winthrop Professor of Genetics at Harvard Medical School and Professor of Health Sciences and Technology at Harvard and MIT, and was a founding member of the Wyss Institute for Biologically Inspired Engineering at Harvard.
He has co-author of over 500 academic papers, 143 patents and co-founded 22 biotech companies. In addition, he has participated in technology development, advising most of the major Genetic Sequencing companies, and he has been at the forefront of genetic research since the 1980s when he was involved with setting up the Human Genome Project.
His impact in the world of genetics has been tremendous.
But Prof Church is also something of a maverick. A left-field thinker. A disrupter.
He is a great supporter of open access genome sequencing and shareable human medical data. He is also keen to bring back extinct species, such as the Woolly Mammoth (Click here for more on this idea).
The return of the woolly mammoth. Source: Phys
Most recently, however, his name has been associated with a new company called Nebula Genomics.
Nebula Genomics is a direct-to-consumer DNA sequencing company.
What does that mean?
The company provides analysis of DNA directly to consumers who would like to know whether they have a genetic risk factor (or a mutation in their DNA), which is associated with a particular medical condition.
What exactly is a mutation in DNA?
To explain this, we need to understand a little something about basic genetics.
In almost every cell of your body you have DNA (or Deoxyribonucleic acid). It is a molecule that is composed of two chains which coil around each other, forming the famous double helix shape.
 Source: Pngtree
Source: Pngtree
The information stored in DNA provides the template – or the instructions – for making (and maintaining) a particular organism. All of the necessary details are encoded in that amazing molecule.
How is information stored in DNA?
In the image above, you can see strands reaching between the two chains of the DNA molecule, and it is these numerous strands that contain the information.
These strands are made up of a pair of ‘nucleotides’. These are organic molecules which contain one of four bases – the familiar A, C, T & Gs that make up our DNA code. These nucleotides pairings (called ‘base pairs’) join together in long strings of DNA.
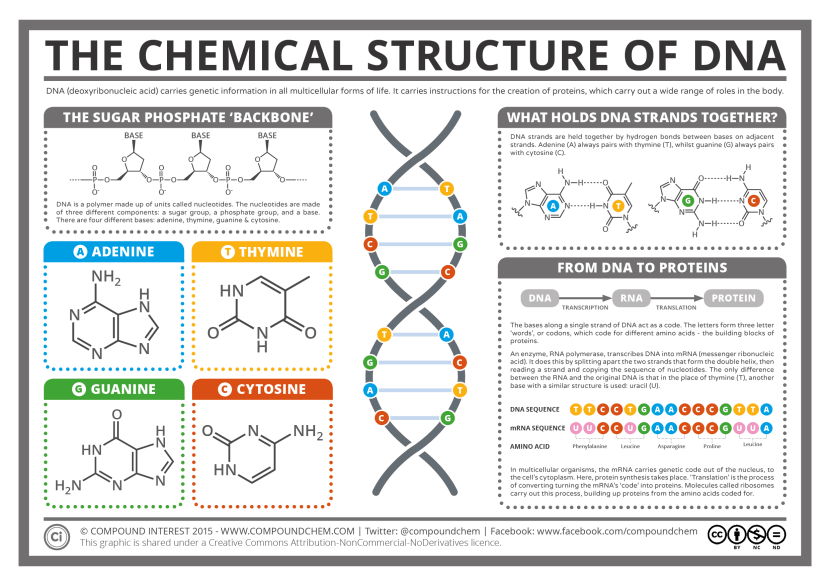 The basics of genetics. Source: CompoundChem
The basics of genetics. Source: CompoundChem
It takes a lot of instructions to make a fully functional human being, and as a result human DNA contains a lot of these base pairs – 3 billion of them. And having so many base pairs results in a lot of DNA in each cell – 2 meters of it. In order to get all that DNA crammed into the nucleus of a tiny little cell, DNA is wound up tightly into thread-like structures called chromosomes.
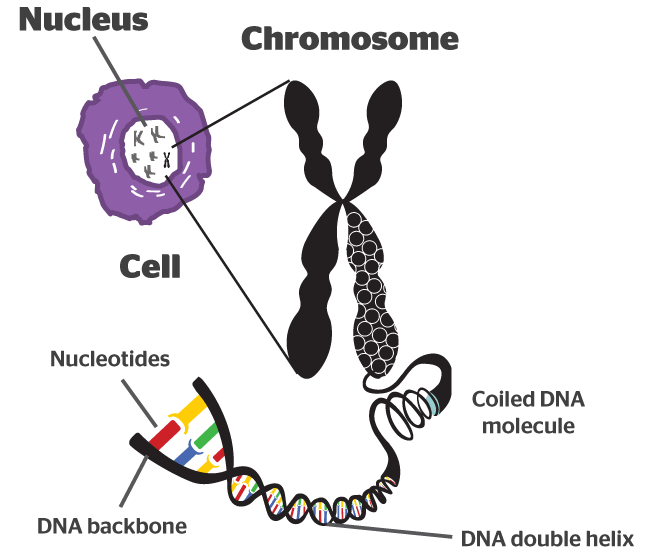 DNA in a chromosome. Source: Byjus
DNA in a chromosome. Source: Byjus
Humans have 23 pairs of chromosomes. And the genetic mutations we are talking about today can occur in any of these chromosomes.
This video does a better job of explaining DNA than I do:
Now, if DNA provides the template for making a human being, it is the small variations in our individual DNA that ultimately makes each of us unique. Every single one of us has small genetic variations.
And these variations come in different flavours: some can simply be a single mismatched base pair (also called a point-mutation or single nucleotide polymorphism (SNP)), while others are more complicated such as repeating copies of multiple base pairs.

Lots of different types of genetic variations. Source: Nature
An example of how these variation can make us who we are is red hair. The fact that an individual has red hair results (in the majority of cases) from a variation in a region of DNA (called MC1R) on chromosome 16.
If that variation is not there: no red hair.
Most of the variants that we have that define who we are, we have had since conception. These are called ‘germ line’ mutations, while those that we pick up during life and that are usually specific to a particular tissue or organ in the body (such as the liver or blood), are called ‘somatic’ mutations.

Somatic vs Germline mutations. Source: AutismScienceFoundation
These variations can occur in regions of DNA that have no apparent use (about 90% of your DNA), but they can also appear in more functional regions (which are called genes). Genes are regions of DNA that provide the instructions for producing proteins, enzymes and pieces of regulatory RNA. These regions provide the instructions (or RNA), which can in some cases be used to make protein. Where genetic mutations fall within these regions, the resulting RNA and protein can carry a variation that will impact their function – sometimes increasing activity, sometimes preventing activity.
An important aspect of these variations in genes is that you have two copies of each gene (called alleles). One copy from your father, and one copy from your mother. Call it mother nature’s insurance policy – if one copy of a gene is faulty, you have a back up. But sometimes one copy is not enough, and this can lead to problems.
In conditions that are influenced by genetics, there are two types of inherited mutations:
- A variant has to be provided by both the parents for a condition to develop – this is called an ‘autosomal recessive‘ disease. In this case, both copies (or alleles) will be mutated, resulting in the condition having a higher chance of developing. An example of this is cystic fibrosis (a condition in which sticky mucus builds up in the lungs and digestive system).
- Only one copy of the variant needs to be provided by one of the parents for a condition to develop – this is called an ‘autosomal dominant’ disease. In this case just one allele is required for a condition to have a higher chance of developing, and a example of this is the neurodegenerative condition of Huntington’s.
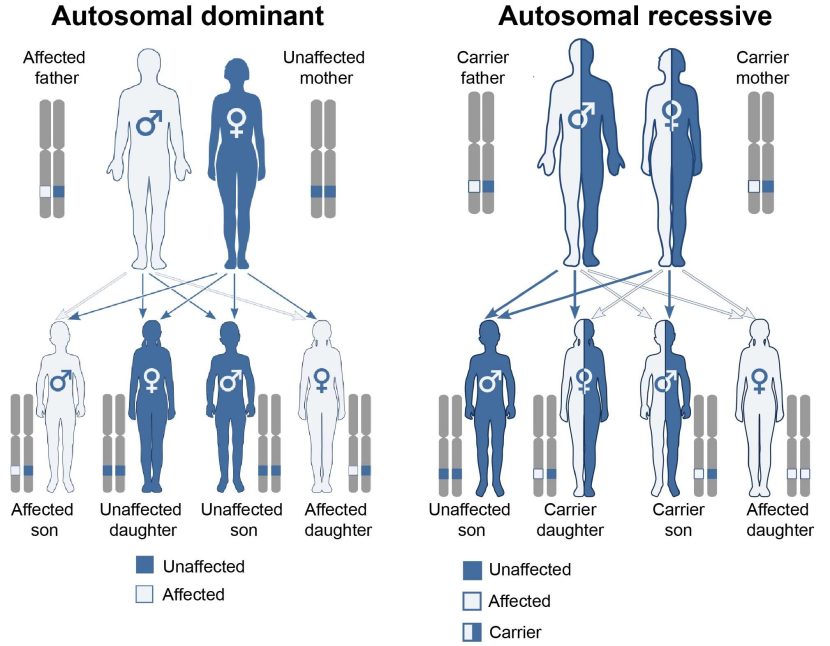
Autosomal dominant vs recessive. Source: Wikipedia
It is important to understand that most of these tiny genetic variation result in no impact on an organism. The region of DNA is not important, or biology is able to adapt and find a way around the problem.
Other variants can infer traits that may be considered beneficial, for example, some people have a genetic mutation in the CCR5 gene. HIV uses that protein to enter into human cells. Thus, if a person has a mutation in CCR5, they are extremely unlikely to become infected by the virus.
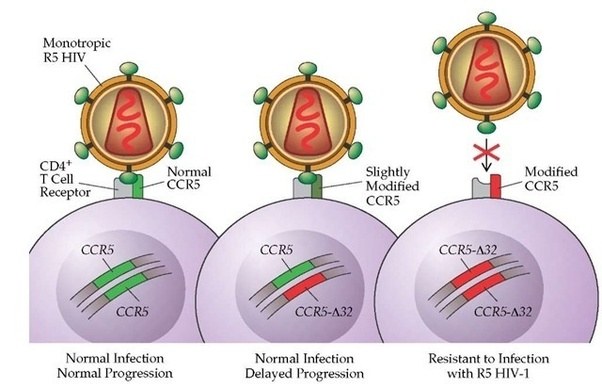 Source: Quora
Source: Quora
There are, however, some genetic variation that are of a more serious nature – leaving us potentially vulnerable to developing a condition (such as BRCA1 mutations in breast cancer). And for a long time, researchers have been interested in identifying genetic variations that are associated with Parkinson’s.
EDITOR’S NOTE: Context is critical in all of these situations, as we have previously discussed in a post on the evolutionary advantage of Parkinson’s – Click here to read that post.
This is very interesting, but is there a genetic association with Parkinson’s?
Since it was first described by good old JP in 1817, there have been suspicions and theories of a hereditary/familial link in Parkinson’s. The condition appeared to be passed on in families – with uncles, fathers, etc also having the condition or symptoms of it.
These patterns have resulted in the idea that there could be a genetic link in Parkinson’s.
It was not until 1997, however, that the first genetic risk factor of Parkinson’s was discovered:

Title: Mutation in the alpha-synuclein gene identified in families with Parkinson’s disease.
Authors: Polymeropoulos MH, Lavedan C, Leroy E, Ide SE, Dehejia A, Dutra A, Pike B, Root H, Rubenstein J, Boyer R, Stenroos ES, Chandrasekharappa S, Athanassiadou A, Papapetropoulos T, Johnson WG, Lazzarini AM, Duvoisin RC, Di Iorio G, Golbe LI, Nussbaum RL.
Journal: Science. 1997 Jun 27;276(5321):2045-7.
PMID: 9197268
In this critical study, the researchers had encountered several independent Italian and Greek families whose members exhibited a similar early onset form of Parkinson’s. The investigators found that the affected members of those families also had genetic mutations in the long arm of chromosome 4.
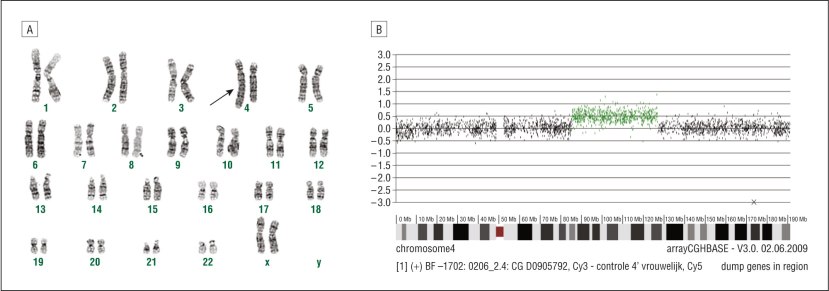
The 23 pairs of chromosomes and variants (green) in chromosome 4. Source: JAMA
It was around this same time that parts of chromosome 4 was sequenced and mapped out, and as a result the researchers were able to determine that the area covering the Parkinson’s associated genetic mutations lay in a gene called SNCA, which provides the instructions for making a protein that we here at the SoPD are rather familiar with: Alpha synuclein.
You can read more about the first Parkinson’s genetic mutations in a previous SoPD post (Click here to read that post)
Over the next few years, more genetic variations that inferred a higher risk of developing Parkinson’s were identified. These discoveries gradually resulted in a list – called the PARK genes – of 20+ regions of DNA where variations were considered risk factors for developing Parkinson’s.
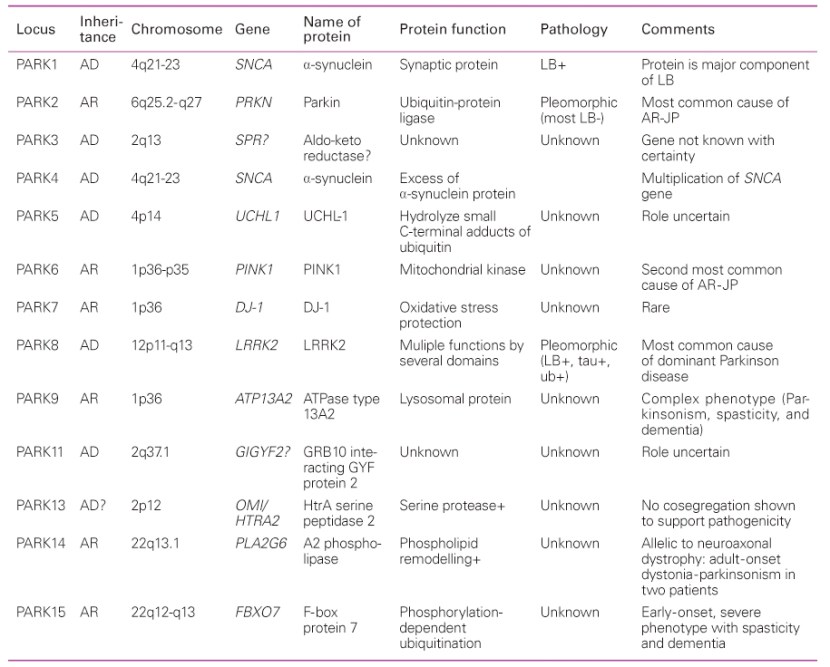
An early list of the PARK genes. Source: JKMA
More recently, a lot of new genetic risk factors for Parkinson’s have been identified. A good example of this is a study published last year (2017):
 Title: A meta-analysis of genome-wide association studies identifies 17 new Parkinson’s disease risk loci.
Title: A meta-analysis of genome-wide association studies identifies 17 new Parkinson’s disease risk loci.
Authors: Chang D, Nalls MA, Hallgrímsdóttir IB, Hunkapiller J, van der Brug M, Cai F; International Parkinson’s Disease Genomics Consortium; 23andMe Research Team, Kerchner GA, Ayalon G, Bingol B, Sheng M, Hinds D, Behrens TW, Singleton AB, Bhangale TR, Graham RR.
Journal: Nature Genet. 2017 Oct;49(10):1511-1516.
PMID: 28892059 (This report is OPEN ACCESS if you would like to read it)
In this study, the researchers conducted a GWAS comparing DNA from 6,476 people with Parkinson’s with DNA from 302,042 controls. They then compared those results with another GWAS dataset from a recent study involving 13,000 people with Parkinson’s and 95,000 controls.
Wait a minute. What is a GWAS?
A genome-wide association study (or GWAS) is an analysis of a set of genetic variants across the entire genome (or all of the DNA in your cells, including mitochondrial), and this analysis is conducted in a large pool of different individuals to see if any variants are associated with a particular trait (or medical condition). It is typically an analysis of single nucleotide polymorphisms (or SNPs; a variation in a single nucleotide). The researcher will check your DNA for the presence of a large set of single nucleotide variations, and then compare them with the results collected from other people.
 Source: Knowgenetics
Source: Knowgenetics
Now as we have discussed above, we all have these tiny mutations. But what a GWAS does, is seek to determine whether people with a particular trait (for example, constrictions in small blood vessels) have a shared single nucleotide polymorphisms, compared to people that do not have that trait. Think of the example of red hair mentioned above, a GWAS analysis of red haired people would point towards a variation on chromosome 16 (in the MC1R gene).
And the result can be presented in what is called a “Manhattan plot” – Manhattan plots get their name because they often look like the city skyline of Manhattan:
 Source: Andrewprokos
Source: Andrewprokos
Across the bottom of a Manhattan plot are the chromosomes, and each dot represents a single nucleotide polymorphisms for that particular region of the DNA. The more dots there are above that region – the more people with a variation in that region – the higher the dot in that particular location. The Manhattan plot in the image below is for an analysis of people with constrictions in small blood vessels, and what you can see is that the number of dark blue dots above chromosome 19 indicate that there are a lot of people who have this trait that also have a genetic variant in that region.
Source: Wikipedia
When researchers who conducted the Parkinson’s study from last year analysed of their data, this was the kind of Manhattan plot they observed:
 Source: PMC
Source: PMC
Immediately you can hopefully see that we are looking at a complicated set of result – there are a lot of ‘buildings’ in this Manhattan city skyline. Lots of genetic variants that could be considered risk factors for Parkinson’s.
The researchers then tested these results in a third data set of DNA from 5,851 people with Parkinson’s and 5,866 controls. From this combined analysis (totalling 26,035 PD cases and 403,190 controls), they found 17 novel risk factors for Parkinson’s (41 in total; 24 had been previously reported).
A great deal of the Parkinson’s-focused biology currently being conducted in research labs around the world is based on the genes (and their functions) identified in these kinds of studies.
Ok, so what is this new company Nebula Genomics doing that is different to other ‘direct-to-consumer’ DNA sequencing companies?
Well, firstly, they appear to be sequencing a lot more of your DNA than some of the other companies currently – those companies generally confine their analysis to specific regions of DNA that are associated with risk of certain medical conditions. In the case of Parkinson’s, these companies will focus on regions such as GAB, LRRK2, SCNA, etc. Nebula is offering to do a much broader analysis.
And the ability of Nebula Genomics to do this is partly a consequence of the very rapidly decreasing cost of sequencing DNA.
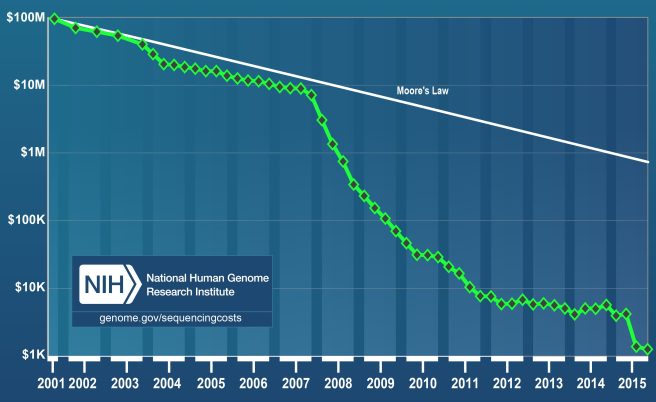 Source: Genome
Source: Genome
Secondly (and more importantly for our conversation here), the company is using a modified business model that differs from many of its competitors.
What do you mean ‘modified business model’?
Traditionally, DNA sequencing was a medical test requested by a clinician who may have suspected a genetic basis to a medical condition in a particular patient. A sample of DNA would be collected from the patient (usually a blood sample) and it would be sent for sequencing and analysis. The results would then be sent back to the clinician who would feed the information back to the patient.
 The good old days. Source: Medicinetoday
The good old days. Source: Medicinetoday
More recently, numerous companies have started selling DNA sequencing services directly to consumers. People sitting at home can request a specimen collection kit (this usually involves a sample of saliva) and sending it to the DNA sequencing company who conduct the analysis. The company then sends the results back to the consumer, and often provides information about resources that can help with interpretation of the results.
 Source: Voices
Source: Voices
This current business model allows the DNA sequencing company to earn money both from the consumer end of their business, but also by providing access to the data to third parties who may be interested in a particular genetic risk factor. The information is annonymised, but recently there have been arrangements that have allowed third parties to reach out to contact individuals of interest.
For example, recently the Pharmaceutical company GlaxoSmithKline and the direct-to-consumer DNA sequencing company, 23andMe signed a $300M equity investment and a four-year research collaboration (Click here to read the press release from GSK).
 Source: Medium
Source: Medium
GSK views this deal as a means of speeding up their clinical trial programme for the development of LRRK2 inhibitors for Parkinson’s – they say as much in some of their recent presentations (Click here to read more about this).
How will this deal help GSK?
GSK has a very active LRRK2-inhibitor programme.
And because some of these genetic variants are extremely rare – even within the Parkinson’s community – they need all the help they can get for recruiting people to take part in their clinical trial. A clinical study of people with LRRK2-G2019S associated Parkinson’s will need to genotype (or DNA sequence) approximately 100 people with Parkinson’s to identify just one individual with a LRRK2-G2019S genetic mutation – that is how rare some of these genetic variants are.
23andMe, however, has more than 10,000 ‘re-contactable’ individuals with Parkinson’s in their database, approximately 3,000 of those ‘re-contactable’ individuals have LRRK2 G2019S carriers. By paying 23andMe for access, GSK can skip a a lengthy recruitment process. Thus, speeding up the clinical trial process.
And GSK/23andMe are not the only companies taking part in such deals. Denali Therapeutics and Centogene recently signed a similar sort of deal (Click here to read the press release).
Source: Twitter
This is good though, right?
Yes, speeding up the clinical trial process is a good thing.
The problem in the current arrangement, however, is privacy. In addition, many individuals who have had their DNA sequenced (without reading the fine print) may not appreciate DNA sequencing companies profitting from their genetic information.
Nebula Genomics is proposing is a new adaptation of this model. They are proposing that if patients pay for their genetic sequencing, they should control who has access to it, AND that they should also be rewarded for that access.
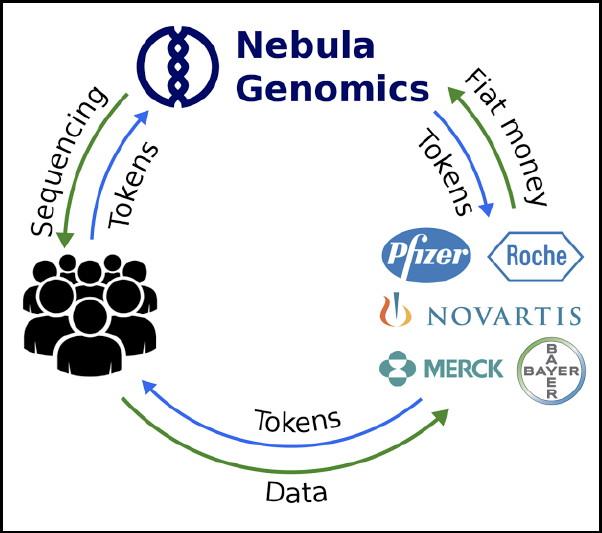 Source: Geneticsandsociety
Source: Geneticsandsociety
The Nebula Genomics business model is based on a system of blockchain protocols.
What is blockchain?
A blockchain is a record-keeping technology. It involves a growing list of records (referred to as blocks) which are linked (or ‘chained’) using cryptography.
How does blockchain work?
Traditionally, when a transactions take place between two individuals, both parties have a record of the transaction. The problem with this situation, however, is that those individual records are stored internally. Since both parties have their own version of the records, there is the potential for fraud or errors to occur.
Blockchain provides a system by which those the records of transactions are maintained on a shared, tamper-proof digital ledger. For a transaction to take place, both parties needs to consent to it.
This video explains how blockchain works:
And here is where there is the potential for individuals to take back control of information related to their DNA with blockchain: the individual will need to give their consent for their genetic data to be offered to third parties, and they could potentially profit from such a situaition.
Such a model places the patients in the middle of the decision making process.
Source: Voices
If big Pharma companies want 50-to-60-year-old males with LRRK2-associated Parkinson’s (and no other notable conditions), they can put out a call and offer to pay individuals the ‘going rate’ for access to their genetic information.
So what does Nebula Genomics offer?
Nebula offers two approaches:
- The consumer pays US$100 and has their DNA sequenced. Then they can either “keep your data private OR continue to receive insights, analytics and rewards on the platform for contributing to the research studies you choose”.
- Consumers can fill in some survey information and refer friends, and be rewarded with tokens that can be used to purchase a ‘low-pass Whole Genome Sequencing’. Alternatively, they can “get matched with research institutions that will cover the cost of your whole genome sequencing” (based on the outcome of their survey results). Under this plan, the individual will receive their DNA results AND get the option to either “keep your data private OR continue to receive insights, analytics and rewards on the platform for contributing to the research studies you choose”.
For people living outside of the US, Nebula Genomics sequencing service will become available by mid 2019.
Click here to learn more about Nebula Genomics.
Is Nebula the first company in this space?
No.
A group called Gene Blockchain was launched in 2013, offering the “World’s First Human Genome Blockchain.”
 Others examples of this business model include Encrypgen , Luna DNA , Longenesis , and Zenome.
Others examples of this business model include Encrypgen , Luna DNA , Longenesis , and Zenome.
This is interesting, but why is genetic information important for the Parkinson’s community?
The obvious use of genetic data is using it to learn more about the biology of Parkinson’s.
Another use of such information is determining who in the Parkinson’s community may respond better to treatments designed to target specific aspects of Parkinson’s, such as the LRRK2 inhibitors we mentioned above OR boosting the waste disposal system – this was the goal of the Ambroxol trial we are currently awaiting the results of. This study was conducted on people with and without genetic mutations in the GBA gene (Click here to read more about GBA).

Ambroxol. Source: Skinflint
It is highly likely that future clinical trials will involve firstly genetically testing participants to determine if they have genetic variants associated with a particular biological function that could be therapeutically targeted. By using platforms like Nebula Genomics, we should hopefully be able to speed up this process by knowing in advance who has which genetic variants.
So what does it all mean?
Increasingly there is a desire from both the affected community and research community to identify genetic variations that are associated with specific medical conditions. Such knowledge can improve our understanding of these ailments and ultimately aid us in better treating them.
For a long time, however, there has been a debate regarding the ownership of genetic information. Now innovative/disruptive new companies are offering an alternative approach for communities to handle their genetic information, and it will be interesting to see how this changes our approach to future research and clinical trials.
By putting the power in the hands of the patients, companies like Nebula are hoping to change the current system and empower the affected community. Feasibility of this new model will rely in large part on “acceptance, adoption, and regulation” of this blockchain approach (or something similar). Regardless of whether it actually succeeds, better availablity of the genetics of Parkinson’s will hopefully help in speeding up the process of clinically testing new therapies for conditions like Parkinson’s.
FULL DISCLOSURE: The information provided here is for educational purposes. The author of this post has had no communication with Nebula Genomics and the information presented here should not be considered an endorsement of Nebula Genomics or the services provided by that company. This post has been written by the author solely for the purpose of sharing the information provided as the author thought the concept behind Nebula Genomics was an interesting idea.
The banner for today’s post was sourced from








I strongly recommend the The PD GENEration study at University of Indianapolis. The test is simple and free. A DNA councillor is assigned to each participant.
After reading your articles I signed up for the most expensive test with Nebula and it was near to useless. The presentation of the results are misleading at best and impractical at worst. You basically are told what your risk is relative to other Nebula participants. NOT relative to a comparable population cohort. And instead o naming known Parkinson associated genes that can have mutations it just dumps out a raw list of SNPs. You then have to take those and manually enter them in a some sort of data cooker to get a little more of a handle but it still does not use nomenclature that is recognizable from the literature. So you manually have to map to the GBA, LRRK2 etc.
The Nebula test was not cheap. I paid almost $1000!!! In contrast the PD GENEration is free and VERY user friendly. I hope you will consider recommending this over Nebula.
Overall, thank you for a wonderful and Herculean effort to make Parkinson research accessible and for a very comprehensive coverage. Your blog has been incredibly helpful and serves a a constant checkpoint. Your work is incredible.
LikeLike