
Very interesting results published last week regarding the bacteria in the intestinal system of people with Parkinson’s disease.
This is an important piece of research because the gut is increasingly being seen as one of the potential start sites for Parkinson’s disease.
In today’s post we will review the results and discuss what they mean.
Bacteria in the gut. Source: Huffington Post
Before you go to bed tonight, contemplate this:
The human gut hosts tens of trillions of microorganisms, including at least 1000 species of bacteria (which is a guess-timate as we are not really sure how many species there are).
And whenever you feel like you are all alone, know that you are not.
You are never alone: tens of trillions of microorganisms are with you!
And there is sooooooo many of these microorganisms, that they can make up as much as 2 kg of your total weight.
What do the microorganisms do?
Ours bodies are made up of microbiota – that is, collections of microbes or microorganisms inhabiting particular environments (or region of our body) and creating “mini-ecosystems”. And whether you like this idea or not, you need them.
The microorganisms in the human gut, for example, perform all manner of tasks for you to make your life easier. From helping to break down food, to aiding with the production of some vitamins (in particular B and K).
That’s great, but what does the bacteria in our gut have to do with Parkinson’s disease?
People with Parkinson’s disease quite often have issues associated with the gastrointestinal tract (or the gut), such as constipation for example. Some people believe that some of these gut related symptoms may actually pre-date a diagnosis of Parkinson’s disease, which has led many researchers to speculate as to whether the gut could be a starting point for the condition.
We have previously discussed the gut and Parkinson’s disease in several posts (click here, here and here to read them).
Today we re-address this topic because a group of scientists from the USA have determined that the populations of bacteria in the guts of people with Parkinson’s disease are different to those of healthy individuals.
Sounds interesting. What exactly is the difference?
Well, before we discuss that, we need a little bit of background.
In 2015, a group of scientists from Finland, published this research paper:

Title: Gut microbiota are related to Parkinson’s disease and clinical phenotype.
Authors: Scheperjans F, Aho V, Pereira PA, Koskinen K, Paulin L, Pekkonen E, Haapaniemi E, Kaakkola S, Eerola-Rautio J, Pohja M, Kinnunen E, Murros K, Auvinen P.
Journal: Mov Disord. 2015 Mar;30(3):350-8.
PMID: 25476529
In this study the researchers compared the fecal microbiomes of 72 people with Parkinson’s disease and 72 control subjects by sequencing the V1-V3 regions of the bacterial 16S ribosomal RNA gene.
Hang on a minute. What does… any of that mean?
Yeah. Ok, that was a bit technical.
The microbiome refers to the genetics of the microorganisms – that is their genomes (or DNA). When researchers want to look at the microbiome of your gut, they do so by collecting fecal samples (delightful job, huh?).
Interesting facts: Fresh feces is made up of approx. 75% water. Of the remaining solid fraction, 84–93% is organic solids. These organic solids consist of: 25–55% gut bacterial matter, 2–25% protein, 25% carbohydrates, and 2–15% fat (Source: Wikipedia).
Still with me?
After collecting the fecal samples, researchers will extract the DNA from the gut bacterial material, which they can then analyse.
And what are the V1-V3 regions of the bacterial 16S ribosomal RNA gene?
The 16S ribosomal RNA gene is universal in bacteria – it is present in all of their genomes/DNA. The genetic sequence of this particular gene is approximately 1,550 base pairs long, and contains regions that are highly conserved (that is they are shared between species) and highly variable (very different between species).
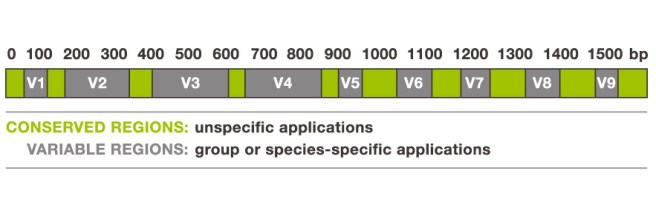

The 16S ribosomal RNA gene. Source: Alimetrics
The gene contains nine of these highly variable regions (called V1 – V9) that display considerable differences in the genetic sequence between different groupings of bacteria. The V2 and V3 regions are considered the most suitable for distinguishing all bacterial species to the genus level (‘genus‘ being a method of classification).
Now scientist can amplify the 16S ribosomal RNA gene by making lots of copies of the highly conserved regions (using PCR) which are shared between bacteria, but then they will genetic sequence the variable sections in between (in this case V2 & V3), which will allow them to discriminate and quantify the different species of microorganisms (such as bacteria) within a particular sample.
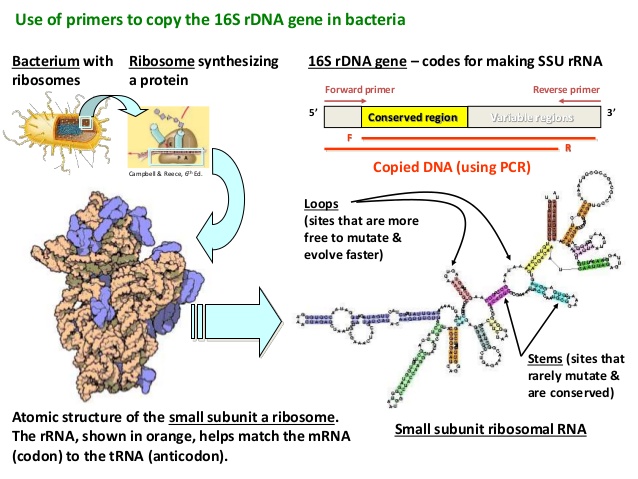
16S rRNA gene analysis – looks complicated. Source: Slideshare
And this is what the scientists in this study did.
They took fecal samples of 72 people with Parkinson’s disease and 72 control subjects, amplified the V1-V3 regions of the bacterial 16S ribosomal RNA gene, and then sequenced the variables regions in between to determine what sorts of bacteria were present (and/or different) in the guts of people with Parkinson’s disease.
The researchers found that there was a reduced abundance of Prevotellaceae in the guts of people with Parkinson’s disease (Prevotellaceae are commonly found in the gastric system of people who maintain a diet low in animal fats and high in carbohydrates, for example vegetarians).
Prevotella multisaccharivorax which belongs to the Prevotellaceae family. Source: MindsofMalady
In addition, the investigators also reported a positive association between the abundance of Enterobacteriaceae and postural instability and gait difficulty symptoms – that is to say, people with Parkinson’s disease who also had postural instability and gait difficulties had significantly more Enterobacteriaceae in their guts than people with Parkinson’s disease who were more tremor dominant.
Due to the design of the study, the researchers were not able to make conclusions about causality from their study. Neither could they tell whether the microbiome changes were present before the onset of Parkinson’s disease or whether they simply developed afterwards. All they could really say was at the time of analysis, they did see a difference in the gut microbiota between people with and without Parkinson’s disease.
And while these same researchers are currently conducting a two year follow up study to determine the stability of these differences over time in the same subjects, they admit that much larger prospective studies are required to address such issues as causality.
Which brings us to the new research published last week:

Title: Parkinson’s disease and Parkinson’s disease medications have distinct signatures of the gut microbiome.
Authors: Hill-Burns EM, Debelius JW, Morton JT, Wissemann WT, Lewis MR, Wallen ZD, Peddada SD, Factor SA, Molho E, Zabetian CP, Knight R, Payami H.
Journal: Mov Disord. 2017 Feb 14. [Epub ahead of print]
PMID: 28195358
The researchers in this study (completely independent from the previous study) applied the same study design as the previous study, but on a much larger scale:
They took samples from a total of 197 people with Parkinson’s disease and 130 healthy controls. And importantly, none of the individual subjects in the study were related (this was an attempt to reduce the effect of shared microbiota between people who live together). Participants were enrolled from the NeuroGenetics Research Consortium in the cities of Seattle (Washington), Atlanta (Georgia) and Albany (New York).
So what did they find?
The researcher’s data revealed alterations in at least 7 families of bacteria: Bifidobacteriaceae, Christensenellaceae, Tissierellaceae, Lachnospiraceae, Lactobacillaceae, Pasteurellaceae, and Verrucomicrobiaceae families
Of particular interest was their observation of reduced levels of Lachnospiraceae in Parkinson’s disease subjects. Lachnospiraceae is involved with the production of short chain fatty acids (SCFA) in the gut. Depletion of SCFA has been implicated in the pathogenesis of Parkinson’s disease (Click here for more on this), and it could potentially explain the inflammation and microglial cell activation observed in the brain (Click here for more on this).
Importantly, they did not replicate the association of Parkinson’s disease with Prevotellaceae (see the previous study above).
The investigators also looked at the medication that the subjects were taking and they found a significant difference in the gut microbiome in relation to treatment with COMT inhibitors and anticholinergics. The effects of COMT inhibitors and anticholinergics on hte microbiome was independent of the effect that Parkinson’s disease was having.
The investigators concluded that Parkinson’s disease is accompanied by ‘dysbiosis of gut microbiome’ (that is, microbial imbalance). Again they could not determine whether the ‘chicken came before the egg’ so to speak, but it will be interesting to see what follow up work in this study highlights.
What does it all mean?
The studies that we have reviewed today provide us with evidence that the bacteria in the guts of people with Parkinson’s disease are different to that of healthy control subjects. Whether the differences between the studies results are due to regional effects (Finland vs USA) will require further investigation. But given that so much attention is now focused on the role of the gut in Parkinson’s disease, it is interesting that there are differences in the gut microbiome between people with and without Parkinson’s disease.
One issue that both studies do not address is whether this difference is specific to Parkinson’s disease and not other neurodegenerative conditions. That is to say, it would have been very interesting if the investigators had included a small set of samples from people with Alzheimer’s disease, for example. This would indicate which differences are specific to Parkinson’s disease as opposed to differences that a general to individuals who have a neurodegenerative condition. If they can tease out medication-related differences (in the second study), then this should be a do-able addition to any future studies.
One would also hope that the researchers will go back and dig a little deeper with future analyses. Using 16S ribosomal RNA gene analysis to determine and quantify the different families of bacteria is analogous to dividing people according to hair and eye colour. The bacteria of our gut is a lot more complicated than this review has suggested. For example, future studies and follow up research could include some genetic techniques that go beyond simply sequencing the 16S ribosomal RNA gene. The investigators could sequence the entire genomes of these species of bacteria to see if genetic mutations within a particular family of bacteria is present in people with Parkinson’s disease.
Easy to say of course. A lot of work, in practise.
There is most likely going to be more of a focus on the gastrointestinal tract in Parkinson’s disease research as a result of these studies. It will be interesting to see where this research leads.
The banner for today’s post was sourced from Youtube
Interesting information. Wishing there would be more research made in developing countries, like Latino american ones.
LikeLike
Hi Fundacion Parkinson Carabobo,
Thanks for your message.
We fully agree! Too little is known are Parkinson’s disease in the developing countries. There does seem to be a growing appreciation of this situation however. For example, it was encouraging to see a good turn out for the recent 1st Pan American Parkinson’s Disease and Movement Disorders Congress in Miami (24th-26th February; http://www.pascongress2017.org/MDS-PAS-2017.htm ). While there were a lot of US based attendees, it was very interesting to note the number of research presenting attendees from Latin american countries (see the program here: http://www.pascongress2017.org/MDS-PAS-2017-Files/PDFs/PASmiami-FP-v9.pdf). So there is research being done in these regions, but certainly not enough. More is required.
We like your website ( http://parkinsoncarabobo.org.ve/) and applaud your efforts to raise awareness about Parkinson’s disease in Venezuela. Keep up the very good work.
Simon
LikeLike
We just completed sequencing the entire genomes of the species found in the faeces of PD patients and healthy controls: https://genomemedicine.biomedcentral.com/articles/10.1186/s13073-017-0428-y
We found increased Verrucomicrobiaceae (Akkermansia muciniphila) and unclassified Firmicutes, whereas
Prevotellaceae (Prevotella copri) and Erysipelotrichaceae (Eubacterium biforme) were markedly lowered in PD
samples. The observed differences reliably separated PD from control with a ROC-AUC of 0.84. Functional
analyses of the metagenomes revealed differences in microbiota metabolism in PD involving the ẞ-glucuronate
and tryptophan metabolism. While the abundances of prophages and plasmids did not differ between PD and
controls, total virus abundance was decreased in PD participants. Based on our analyses, the intake of either a MAO inhibitor, amantadine, or a dopamine agonist (which in summary relates to 90% of PD patients) had no overall influence on taxa abundance or microbial functions.
Our data reveale differences of colonic microbiota and of microbiota metabolism between PD
patients and controls at an unprecedented detail not achievable through 16S sequencing. The findings point to a
yet unappreciated aspect of PD, possibly involving the intestinal barrier function and immune function in PD
patients.
LikeLike
Hi Prof Wullner,
Many thanks for your message. We have actually looked at your excellent study soon after it was published in a follow up to this post (that post can be found here: https://scienceofparkinsons.com/2017/05/03/trying-to-digest-gut-research/). The study is very interesting – certainly a big step forward on the previous 16S studies – and the results were fascinating and encouraging, especially the reduction in Prevotellaceae as this seems to be the one consistent trend between the results from other groups.
I was particularly intrigued, however, by the decrease in total virus abundance you found in the Parkinsonian participants. Not only did this result demonstrate the depth of your studies analysis, but it also baffled me as I had been assuming that there would be either no difference or perhaps a higher level of viral abundance. I would not have bet on a reduction in abundance. Are you following up that aspect of the research?
Kind regards,
Simon
LikeLike
Hey Simon.
My Dad has Parkinson’s and I’ve been doing a ton of research recently. I’ve always been interested in the gut-brain link and bacteria,it’s kind of a hobby of mine. I apologise for my laymen’s descriptions, I am by no means a scientist but call it a “gut feeling” if you will.
I found it interesting that there seems to be a link between quitting smoking and Parkinson’s onset and how smoking seems to, in some way, slow the progress of Parkinson’s. As crazy as that may sound.
I did a little digging and found some articles on the impact of smoking on gut flora and how this changes when you quit smoking and there seems to be an interesting link between which microbiome landscape while smoking and how quitting smoking changes this to something more like that of Parkinson’s patients.
https://www.ncbi.nlm.nih.gov/pubmed/29626219 – Smoking and the intestinal microbiome
https://www.ncbi.nlm.nih.gov/pubmed/25072500 – Smoking cessation alters intestinal microbiota
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6024871/ – Changes of Colonic Bacterial Composition in Parkinson’s Disease and Other Neurodegenerative Diseases
I can’t help but wonder if these little guys living in our guts don’t hold the secret to understanding why we get Parkinson’s and how to prevent it.
I just thought I share my “gut feeling” with you.
Thanks
LikeLike
Hi Terry,
Thanks for sharing your gut feeling – this is where many of the best ideas come from. Seriously, I get researchers writing to me, saying ‘hey, made an interesting connection with something I read in your comments’. Sorry to hear about your Dad’s diagnosis – I hope the medication is working well for him. I like your proactive attitude with investigating your hunches.
The gut is such a difficult beast to address. There is a universe in there that we know so little about. And unfortunately, there are no common (‘house keeping’) bacteria that everyone in the world shares so it is difficult to conduct comparative analysis. An expert here in Cambridge suggested that we are only aware of about 50% of the ‘little guys’ in there, the rest is a mystery. And the broad families of bacteria analysis we do is akin to comparing redheaded people with black headed people.
All of that said, just because it is a complex system does not mean that we can no investigate it. And I am intrigued with the connection you have made with smoking. I was aware of the smoking-PD angle, but had never considered it from a gut point of view. That is REALLY interesting – may be worth a separate post. Let me have a read of this material and I’ll come back to you.
Kind regards,
Simon
LikeLike