|
# # # # Epigenetics is the study of how non-genetic factors affect genetics. It explores the influence of lifestyle or environmental factors on the activity surrounding our DNA. Recently researchers have been investigating epigenetics within the context of Parkinson’s, and they have discovered a “master switch” that could represent an important target for future therapeutic treatments for the condition. The research focuses on a protein called TET2. In today’s post, we will discuss what epigenetics means, review the new research, and consider what the implications of these new findings could be. # # # # |
 Not the same. Source: AutismBlog.
Not the same. Source: AutismBlog.
Identifical twin studies have been extremely useful to our understanding of conditions like Parkinson’s (Click here to read a previous SoPD post on twins research in Parkinson’s)
But the funny thing about identical twins is that in 100% of cases they aren’t identical.
Yeah sure, they initially share exactly the same DNA, but the further they get away from that magically moment of conception, the less alike they are.
 Same DNA, subtle differences. Source: National Geographic
Same DNA, subtle differences. Source: National Geographic
And we’re not talking about personality or likes/dislikes here.
As biological organisms twins diverge significantly after conception and as the age through life. But if they share the same DNA – the same genetic blueprint – how are these difference possible?
The answer to this question may lie in epigenetics.
What is epigenetics?
Epigenetics (Epi, Greek for ‘over’ or ‘above’ and Genetics,…well, you should be able to work that one out).
Epigenetics is the study of heritable changes (meaning transmissible from parent to offspring) in an organism that are not caused by alteration of the genetic code itself. These variations may result from internal or external factors that cause regions of DNA to turn on and off.

Do you lean left or right? Source: 2ndActHealth
Can you remind me what is a gene?
Good question. Before we discuss epigenetics, let’s understand a little about genetics.
In almost every cell of your body you have DNA (or Deoxyribonucleic acid). It is a molecule that is composed of two chains which coil around each other, forming the famous double helix shape.
 The double helix. Source: Pngtree
The double helix. Source: Pngtree
The information stored in DNA provides the template or the instructions for making (and maintaining) a particular organism. All of the necessary details are encoded in that amazing molecule.
How is information stored in DNA?
In the image above, you can see strands reaching between the two chains of the DNA molecule, and it is these numerous strands that contain the information.
These strands are called ‘nucleotides’. These are organic molecules which contain one of four bases – the familiar A, C, T & Gs that make up our DNA code. These nucleotides form pairs (called ‘base pairs’) which then join together in long strings of DNA.
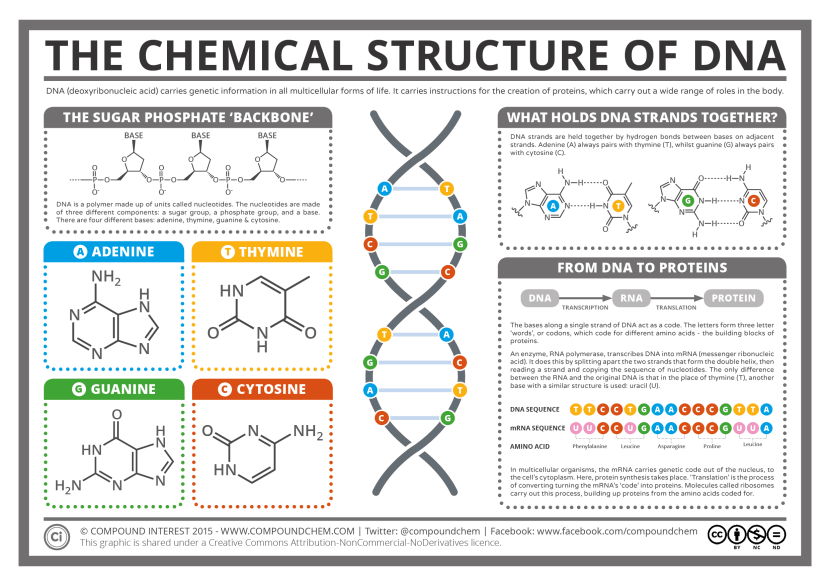
The basics of genetics. Source: CompoundChem
It takes a lot of instructions to make a fully functional human being, and as a result human DNA contains a lot of these base pairs – 3 billion of them. And having so many base pairs results in a lot of DNA in each cell – 2 meters of it. In order to get all that DNA crammed into the nucleus of a tiny little cell, DNA is wound up tightly into thread-like structures called chromosomes.
 DNA in a chromosome. Source: Byjus
DNA in a chromosome. Source: Byjus
Humans have 23 pairs of chromosomes. And the mutations we are talking about today can occur in any of these chromosomes.
This video does a better job of explaining DNA than I do:
All along the long strings of DNA, there are regions that can be ‘read’ and transcribed into RNA – this is the first step in the production of protein. Each of these regions is called a gene. They can provide the instructions for making a particular protein. And we have discovered more recently a new world of RNA that are not used to make protein, but they play a regulatory role in cells. These RNA are called non-coding RNA (ncRNA), and that is a topic for a difference post.
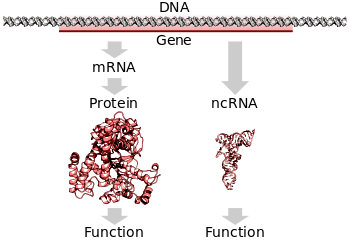 Source: Wikipedia
Source: Wikipedia
It is at the level of the gene that we can start to look at the biological influence of epigenetics.
Epigenetic changes can alter the physical structure of DNA, but it is the activity surrounding the transcribing of genes that really affects biological outcomes in living organisms.
Can you give an example of this kind of epigenetic activity?
A good example of epigenetic activity is DNA methylation.
What is DNA methylation?
DNA methylation is a process by which methyl groups are added to DNA. And before you ask: A methyl group is an alkyl derived from the chemical methane.
 DNA methylation. Source: Labclinics
DNA methylation. Source: Labclinics
When methyl groups are added to DNA, it blocks the DNA from being transcribed. A gene that has been methylated is essentially shut down.
Many of the genes that are important in the early years of development are methylated during the later years of life as they are no longer needed. Interestingly, cells from people with progeria (a rare genetic condition that make affected individuals appear to be older than they are) exhibit DNA methylation patterns that are very similar to those found in normal ageing.
Below is a video of Sam Berns who was diagnosed with Progeria at 2 years of age – his outlook on the world is pretty cool:
If nothing I have written above makes any sense, this video explains provides a useful bit of background information regarding epigenetics:
|
# RECAP #1: Epigenetics is the study of the influence of non-genetic factors (such as lifestyle or diet) on genetics. A good example of epigenetic activty is DNA methylation, where a methyl group is added to DNA and alters the ability for that section of DNA to be transcribed into RNA. # |
Ok, so what does all this epigenetics stuff have to do with Parkinson’s?
There has recently been some interesting research focused on a protein called TET2.
What is TET2?
TET2 is one of the three proteins in the TET (Ten-Eleven Translocation) family – TET1, TET2 and TET3. These proteins are involved in DNA methylation.
Specifically, TET2 promotes DNA demethylation.
Demethylation?
The removal of a methyl group.
DNA methylation = the addition of a methyl group (blocking gene transcription); DNA demethylation = the removal of a methyl group (which has the effect of opening genes back up for business).
DNA demethylation can re-activate genes that may have been methylated perhaps during development. As a result, TET2 appears to play an important role in cellular reprogramming (Click here to read more about this). Such a role could have serious consequences as we shall discuss further below.
Ok, so how is TET2 associated with Parkinson’s?
Two research reports have recently linked this protein with Parkinson’s – the first study hinted at anassociation and the second report provided strong evidence for a connection.
Here is the first study:
 Title: TET2-mediated Cdkn2A DNA hydroxymethylation in midbrain dopaminergic neuron injury of Parkinson’s disease.
Title: TET2-mediated Cdkn2A DNA hydroxymethylation in midbrain dopaminergic neuron injury of Parkinson’s disease.
Authors: Wu TT, Liu T, Li X, Chen YJ, Chen TJ, Zhu XY, Chen JL, Li Q, Liu Y, Feng Y, Wu YC.
Journal: Hum Mol Genet. 2020 May 28;29(8):1239-1252.
PMID: 32037456
In this study, the researchers found high levels of TET2 in models of Parkinson’s.
Specifically, they found that TET2 was significantly increased in mouse dopamine neurons (these are the types of cells that are particularly vulerable in PD in humans), when the cells are exposed to a neurotoxin (MPTP) that is used to model Parkinson’s.
Given this effect, the investigators wanted to determine if blocking the increase in TET2 levels would reduce the dopamine neuron cell death associated with exposure to MPTP. They found that reducing TET2 levels not only protected dopamine neurons, but also rescued the behavioural issues of the MPTP mouse model of Parkinson’s.
And these results have recently been replicated and significantly expanded by a group of researchers at the Van Andel Institute in the USA.
Here is their research report:
 Title: Epigenomic analysis of Parkinson’s disease neurons identifies Tet2 loss as neuroprotective.
Title: Epigenomic analysis of Parkinson’s disease neurons identifies Tet2 loss as neuroprotective.
Authors: Marshall LL, Killinger BA, Ensink E, Li P, Li KX, Cui W, Lubben N, Weiland M, Wang X, Gordevicius J, Coetzee GA, Ma J, Jovinge S, Labrie V.
Journal: Nat Neurosci. 2020 Aug 17. Online ahead of print.
PMID: 32807949 (a preprint of this report is OPEN ACCESS on bioRxiv)
In this study, the researchers examined DNA methylation specifically in the context of Parkinson’s.
They did this by analysing neurons collected from post-mortem brains of 57 people with Parkinson’s (and comparing them with neurons from 48 unaffected/control cases). They found that the Parkinson’s brains had a widespread increase in levels of DNA demethylation.
Has this ever been reported before?
It has.
There have been few epigentic analyses of the Parkinson’s brain, but a study in 2012 reported a lower level of methylation in the PD brain compared to control brains (Click here to read more about this).
But in this new study, however, the investigators reported that this increase in DNA demethylation was associated with high levels of TET2.
And interestingly this increase in TET2 activity also increased with the stage of the disease (based on PD Braak staging). This observation raises the intriguing possibility that epigenetic dysregulation could potentially be contributing to the overall progression of the condition observed in patients with Parkinson’s over time.
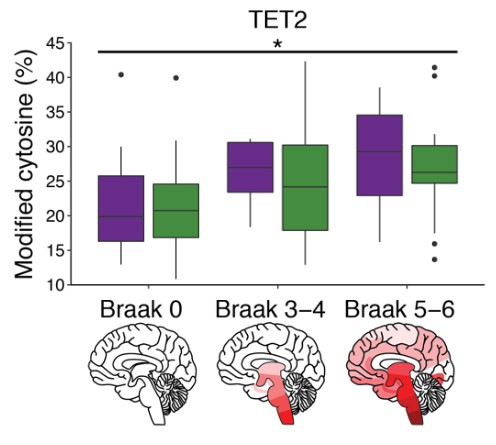 TET2 levels: Females in purple, males in green. Source: BioRxiv
TET2 levels: Females in purple, males in green. Source: BioRxiv
A big difference between the two groups of brains used for this analysis (control brains and PD brains), is that the previous owners of the PD brains had been taking Parkinson’s medication (like L-dopa) regularly before passing away. So the investigators were curious to see if Parkinson’s medication may be a contributing factor to the DNA methylation changes observed in PD, but they found no evidence of this – Parkinson’s medication did not account for the DNA methylation alterations, suggesting that the changes are specific to the condition itself.
When the researchers dug deeper into the data to see what types of genes were affected, they found there was a convergance on genes involved in cell cycle signaling (think cell division) and immune activation (think inflammation).
This is interesting for several reasons – first, activation of cell cycle genes has been implicated in the death of dopamine neurons before (Click here for an example of this), and second, TET2 has recently been reported to regulate neuroninflammation:
 Title: TET2 Regulates the Neuroinflammatory Response in Microglia.
Title: TET2 Regulates the Neuroinflammatory Response in Microglia.
Authors: Carrillo-Jimenez A, Deniz Ö, Niklison-Chirou MV, Ruiz R, Bezerra-Salomão K, Stratoulias V, Amouroux R, Yip PK, Vilalta A, Cheray M, Scott-Egerton AM, Rivas E, Tayara K, García-Domínguez I, Garcia-Revilla J, Fernandez-Martin JC, Espinosa-Oliva AM, Shen X, St George-Hyslop P, Brown GC, Hajkova P, Joseph B, Venero JL, Branco MR, Burguillos MA.
Journal: Cell Rep. 2019 Oct 15;29(3):697-713
PMID: 31618637 (This report is OPEN ACCESS if you would like to read it)
In this study, the researchers reported that TET2 regulated a pro-inflammatory response in microglial cells of mice injected with an inflammation-inducing chemical (LPS). Microglia are the resident immune cells in the brain. They also reported that in both postmortem Allzheimer’s brains and the brains of Alzheimer’s mice, microglia displayed high levels of TET2. When they reduced TET2 in the microglia of the Alzheimer’s mice
This association with inflammation made the Parkinson’s researchers question if TET2 might be involved in the immune response in Parkinson’s, and so they tested this by inactivating TET2 in an inflammatory model (induced by LPS) of Parkinson’s. The investigators found that reducing TET2 fully prevented dopamine neuron cell loss in those mice and reduced inflammation.
The researchers proposed that epigenetic dysregulation may result in raised levels of TET2 in people with Parkinson’s, which may “lead to a compromised neuronal system that has greater vulnerability to inflammation-induced neurotoxicity“.
 The researchers concluded: Based on our study, reducing TET2 in patients with PD may be a new therapeutic intervention that could dampen neuroinflammatory responses and protect against neurodegeneration”.
The researchers concluded: Based on our study, reducing TET2 in patients with PD may be a new therapeutic intervention that could dampen neuroinflammatory responses and protect against neurodegeneration”.
|
# RECAP #2: TET2 is a protein that promotes DNA demethylation, allowing for the re-activation of a methylated gene. If uncontrolled, DNA demethylation may result in inappropriate genes being re-activated which may have consequences. Researchers have found that cells from Parkinsonian brains have high levels of TET2, and this is associated with activation of cell cycle and inflammatory genes. Reducing levels of TET2 is neuroprotective in models of Parkinson’s. # |
Very interesting. Are there any TET2 inhibitors available for me to try?
No there are not.
The senior scientist on the second TET2 report discussed in this post, Dr Viviane Labrie of the Van Andel Institute in Grand Rapids (Michigan), was quoted as saying that:
“Temporarily reducing TET2 activity could be one way to interfere with multiple contributors to the disease, especially inflammatory events, and protect the brain from loss of dopamine-producing cells”. This could “dampen an overactive immune system and protect against neurodegeneration” (Source).
But manipulating levels of epigenetic proteins (especially TET2) is a tricky business.
Why?
TET2 is a widely recognised as a tumor-suppressor gene – meaning a gene that prevents cancers from occurring. It is altered in ~4% of all cancers (over 30% in many leukemias), and its deletion is sufficient to cause myeloid and lymphoid malignancies in mice (Click here to read more about this).
Thus, we are going to have to be careful in how we approach in how we go about reducing TET2. This is not to say that it can not be done – there are more than 7 epigenetic treatments approved for use in oncology (Click here to read more about this) – but it is tricky.
Perhaps a better initial approach may be to reduce the downstream inflammation before dealing with the epigenetic dysregulation (Click here to read a recent SoPD post on inflammation in PD).
So what does it all mean?
Epigenetics is the study of heritable changes that do not involve alterations in the DNA sequence. These changes can be brought on by environmental or lifestyle factors. Recently researchers have reported that epigenetic changes may be playing a role in Parkinson’s – even potentially driving the progression of the condition.
Efforts to pharmacologically manipulate epigenetic activity in neurological conditions like Parkinson’s should be further investigated, but also considered rather blue sky by readers of this blog. Regulating epigenetic components is an extremely complicated game to play.
While the discovery of epigenetic dysfunction in the Parkinsonian brain is very exciting, a crucial immediate question to be addressed is how these epigenetic changes are coming about? What causal factors might be driving them?
And another important question is how to address this crucial immediate question?
# # # # #
Before we finish – I hate to end a post on a sad note, but this is necessary.
Here at the SoPD we were really gutted to learn of the passing of Dr Viviane Labrie of the Van Andel Institute (Press release). She was the lead senior scientist on the second TET2 report discussed in this post. A rising star in the field of the Parkinson’s research, Viviane seemed to have a stellar career ahead of her.
Her passing is a tragic loss.
 Dr Viviane Labrie ( 1980-2020). Source: VanAndel
Dr Viviane Labrie ( 1980-2020). Source: VanAndel

All of the material on this website is licensed under a
Creative Commons Attribution 4.0 International License
You can do whatever you like with it!
The banner for today’s post was sourced from MHH

Epigenetics has fascinated me ever since I discovered that my sensitivity to dental mercury might have been caused by the experiences of a great-grandfather I never knew who died of “hatter’s disease” due to making hats with mercury compounds in NYC, early in the previous century.
I am inclined to agree with you that downstream modulation of the inflammatory process is more predictable and controllable than suppressing a protein (TET2) that removes methyl groups that are blocking various genes, only some of which are producing inflammatory proteins.
Longvida curcumin and baicalin, for example, block the TLR2 and TLR4 receptors on microglia that would otherwise be stimulated by aggregated alpha-synuclein and neuromelanin released from dopamine neurons as they are damaged by immune attacks, and those two supplemennts have a very benign side-effect profile.
Also, if I am reading your diagrams and article correctly, it is not only the case that TET2 has an impact on inflammation; it is also that case that inflammation increases the generation of TET2. And so, even though there is no substance that one can take to reduce levels of TET2 (which, as already noted, is not necessarily a good idea anyway), it seems to me that reducing inflammation (e.g., by non-epigenetic means) would quell the production of TET2 to a certain degree, which could in turn result in reduced inflammation.
And so, it appears to me that TET2 is yet another factor in a set of inter-twined feedback loops that could be broadly characterized as part of the autoimmune process that eats dopamine neurons in the course of Parkinson’s progression.
We already know about the loop in which damaged neurons spill their guts into the inter-cellular space, stimulating microglial receptors so that microglia switch into classical (M1) activation, generating cytokines which trigger apoptosis in bystander neurons, starting their decline and thus re-igniting another cycle of autoimmune inflammation.
To that, it seems that we can now add a process by which inflammation triggers an increase in TET2 production (by whatever means), which in turn starts to remove methyl groups from genes that, once unblocked, start to copy their proinflammatory proteins, which cause more inflammation, re-triggering this epigenetically-grounded inflammatory cycle, while also feeding into the first kind of inflammatory loop that directly involves the death of neurons.
So this could be yet another piece of the puzzle in deconstructing the complex of interacting processes that implement the inflammatory furnace that consumes dopamine neurons over the course of the illness.
LikeLike
Also, I’m wondering: Do we know which genes are being unblocked by TET2’s demethylation, what proteins those unblocked genes are producing, and what receptors on microglia are being stimulated by the increased presence of those proteins? If we knew that, then we might devise downstream methods for blocking the receptors for those proteins, which would providing yet another means for damping down the inflammatory process.
LikeLike
OK, there’s something I’m not sure of: Is the effect of TET2 on microglia a direct effect (stimulating an immune response themselves), or it is an indirect effect that involves TET2 removing methyl groups from *genes* that then produce increased levels of *their* proteins, that then stimulate a microglial response?
LikeLike