|
One of the most common observations that people make when they attend a Parkinson’s disease support group meeting is the huge variety of symptoms between sufferers. Some people affected by this condition are more tremor dominant, while others have more pronounced gait (or walking) issues. In addition, some people have an early onset version, while others has a very later onset. What could explain this wide range of features? A group of Stanford researchers have recently proposed an interesting new idea regarding our understanding of genetics that could partly explain some of this variability. In todays post I speculate on whether their idea could be applied to Parkinson’s disease. |

Source: Discover
Earlier this year an interesting study was published in the prestigious journal Nature on the topic of the genetics of height (yes height. Trust me, I’m going somewhere with this):

Title: Rare and low-frequency coding variants alter human adult height
Authors: Marouli E, Graff M, Medina-Gomez C, Lo KS, Wood AR, Kjaer TR, Fine RS, Lu Y, Schurmann C,………at least 200 additional authors have been deleted here in order to save some space…….EPIC-InterAct Consortium; CHD Exome+ Consortium; ExomeBP Consortium; T2D-Genes Consortium; GoT2D Genes Consortium; Global Lipids Genetics Consortium; ReproGen Consortium; MAGIC Investigators, Rotter JI, Boehnke M, Kathiresan S, McCarthy MI, Willer CJ, Stefansson K, Borecki IB, Liu DJ, North KE, Heard-Costa NL, Pers TH, Lindgren CM, Oxvig C, Kutalik Z, Rivadeneira F, Loos RJ, Frayling TM, Hirschhorn JN, Deloukas P, Lettre G.
Journal: Nature. 2017 Feb 9;542(7640):186-190.
PMID: 28146470
In this study, the researchers – who are part of the GIANT consortium – were analysing DNA collected from over 700,000 people and trying to determine what genetic differences could influence height.
Height is not important for music. Source: Imgur
Why study height?
Good question. There are several reasons:
Firstly, it is easy to accurately measure. Second, the researchers believed that if we can master the complex genetics of something simple like height maybe what we learn will give us a blueprint for how we should study more complex medical disorders that have thus far eluded our complete understanding.
Ok, so what did they find?
Well….
This new research report was a follow up study to a previous report published in 2014. In that previous report, the researchers had used DNA from 253,288 individuals and discovered 697 genetic variations (alterations in the DNA) that affect height (Click here to read an OPEN ACCESS version of that report). Each of those genetic variants had only a tiny impact on the height of an individual (increasing it by mere millimetres), and collectively they could only explain about 16 percent of the variation in height (for people of European ancestry at least).
In their latest report, the same investigators reported that they have now analysed DNA collected from over 700,000 people and that they had found 83 additional genetic variants. Interestingly, 24 of these newly discovered genetic variants affect height by more than 1 cm (4/10 of an inch), but they are extremely rare genetic differences that only becoming apparent in this larger dataset.
And with these new findings, approximately 27.4 percent of the heritability of height can now be accounted for by genetics.
Which begs the question you are probably pondering….
What about the other 72.6%?
Great question.
And the answer is really simple: we don’t know.
Previous estimations based on family and twin studies have suggested that genetics accounts for at least 80% of height (Click here to read more about this). Thus, we are missing a huge chunk of the picture.
What other evidence is there for “none genetic” influences on height?
Global trends in height certainly suggest that other factors can affect height.
This is the largest longitudinal study of height:

Title: A century of trends in adult human height.
Authors: NCD Risk Factor Collaboration (NCD-RisC).
Journal: Elife. 2016 Jul 26;5. pii: e13410.
PMID: 27458798 (This article is OPEN ACCESS if you would like to read it)
In this study, the investigators analysed 1472 population-based studies, with measurements of height taken from 18.6 million participants to estimate mean height for people born between 1896 and 1996 in 200 countries (it is a vast and truly amazing study!).

Trends in European country height. Source: Elife
Remarkably, over the last hundred years, South Korean women and Iranian men have on average become 20.2 cm and 16.5 cm taller, respectively. Such a huge jump in height could not possibly be due solely to genetics.
Could diet or improved quality of life being having an effect?
Researchers in this field have found that foetal growth (itself related to maternal size, nutrition and environmental exposures), general nutrition, and also infections during childhood and adolescence are important determinants of eventual height. But, they only account for a small difference.
In addition, changes in the overall quality of life do not always account for the difference. An impressive rise in the overall height of the population of Japan stopped in people born after the early 1960s, even though their overall quality of life increased significantly with economic growth during that period. And as a result South Koreans are now taller than their Japanese neighbours.
I guess the take home message is, we really don’t understand what determines our height.
Interesting. Really. But what does any of this got to do with Parkinson’s disease???
Indeed. To business.
A group of researchers at Stanford University have proposed an idea that could help in understanding the “72.6%” (the unknown percentage from the GIANT study mentioned at the top of this post) and it may actually alter the way we look at genetics. Plus, (and I am speculating here) it could also help us in understanding the variability in symptoms between individuals with Parkinson’s disease.
They have recently published this perspective of their theory:

Title: An Expanded View of Complex Traits: From Polygenic to Omnigenic.
Authors: Boyle EA, Li YI, Pritchard JK.
Journal: Cell. 2017 Jun 15;169(7):1177-1186.
PMID: 28622505
In their write up, the researchers proposed three primary ideas:
- A small number of “core genes” “provide mechanistic insights into disease biology and may suggest druggable targets.” These are genes that harbour common genetic variations with obvious clinical or biological effects. Think of the gene SNCA which encodes the alpha synuclein protein. Mutations in this gene are known to increase ones risk of developing Parkinson’s disease. It is a well known (or “core”) gene associated with Parkinson’s disease.
- Next, there are a larger number of “peripheral genes”. These are believed to drive a particular disease through cellular networks in specific cell types.
- Thirdly, there is an even larger number of minor peripheral genes and disease-associated variants that contribute a “non-zero effect” (meaning that their effect is tiny, but not entirely insignificant). These minor peripheral genes may outnumber core genes by 100:1 or even 1000:1, we really don’t know. But their tiny individual effects could collectively dominate heritability when they are all pooled together. The issue with these rare variants, however, is that they are difficult to detect on a population-wide basis and may not be easily categorised based on our current understanding of genetics. It is this new third concept, that is the central piece of what authors have termed “omnigenics”.
The basic idea of their theory is that “most heritability can be explained by effects on genes outside core pathways”. In effect, most of the heritability of traits may not be due to a handful of core genes, but to the tiny contributions from a vast number of peripheral genes that function outside the actual disease pathways. The degrees of separation are therefore not un-influential.
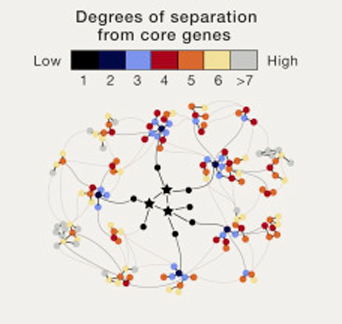
Source: Cell
The authors of the perspective suggested that:
“Intuitively, one might expect disease-causing variants to cluster into key pathways that drive disease etiology. We propose that gene regulatory networks are sufficiently interconnected such that all genes expressed in disease-relevant cells are liable to affect the functions of core disease-related genes and that most heritability can be explained by effects on genes outside core pathways.”
In other words, they suggests we should move away from a simple Polygenic view of inheritance (which involves one characteristic that is controlled by two or more genes), and have a more omnigenic approach (minor peripheral genes influencing variance).
Think back to the height example above, and how a few rare genetic variations had a big impact on height, but they were difficult to find in a small sample population of only 250,000 people. It was not until the researchers expanded their analysis to 700,000 samples that they found them. The bulk of height that we could explain genetically was down to a lot of minor genes having a tiny effect.
And this applies to all heritable traits, not just height.

Source: Boundless
In an effort to test/prove their point, the omnigenic researchers re-analysed the data from the 2014 GIANT study study on height, and they found that for all single nucleotide mutation across the entire genome (the parts of DNA encoding protein), each mutation is associated with an average increase in height of 0.14 mm. Thus, as many as 100,000 single-letter DNA variants can influence a person’s height, at the level of just one-tenth of a millimetre. Not all tall people with have all of these genetic variants, but they may have some of them, which collectively adds to the height of the individual.
The researchers also applied their approach to DNA databases for schizophrenia, rheumatoid arthritis and Crohn’s disease. They found an interesting pattern: minor variants that apparently affect these conditions are present in sections of DNA that are active in the particular cells relevant to the type of disease: neurons for schizophrenia, and immune cells for the two autoimmune diseases.
Ok, but again how does this relate to Parkinson’s disease?
Well, we currently know of 23 genetic variations that infer increased risk of developing Parkinson’s disease (click here to read our page on the genetics of Parkinson’s). These “core” genes are referred to as the PARK genes, with PARK1 being a mutation in the alpha synuclein gene.
In addition to these PARK genes, there are a host of other genes that have been shown to affect risk of developing Parkinson’s disease (for example, mutations in GBA, a gene encoding the enzyme glucocerebrosidase). These could be considered the “peripheral” genes that were discussed above. Collectively, all of these variations account for between 20% of people with Parkinson’s disease (Source). Is this starting to sound a bit like the height study we discussed above?
Perhaps if we apply the idea of omnigenics to Parkinson’s disease we will find that there are hundreds/thousands of minor peripheral genes that are influencing other aspects of the disease, such as age of onset, cognitive decline, and even severity of motor features (tremor, etc). It may in part explain some of the variability observed between members of families affected by Parkinson’s disease (Click here to read more about this), particularly the differences observed in the twins research.
Twins research?

Identical twins. Source: AutismBlog.
In the world of Parkinson’s disease research, twins are fantastic!
I have previously discussed the curious case history of the Gernsheimer twins who have shared the same genetics and environment and yet one of them has Parkinson’s disease (Click here to read that post). But the research looking at twins and Parkinson’s disease really is fascinating.
For example, this study:

Title: Parkinson disease in twins: an etiologic study.
Authors: Tanner CM, Ottman R, Goldman SM, Ellenberg J, Chan P, Mayeux R, Langston JW.
Journal: JAMA. 1999 Jan 27;281(4):341-6.
PMID: 9929087 (This article is OPEN ACCESS if you would like to read it)
In this study, the scientists screened 19,842 male twins enrolled in the National Academy of Sciences/National Research Council World War II Veteran Twins Registry. 163 pairs of twin were identified in which at least 1 twin had developed Parkinson’s disease (and medical records were available).
When a diagnosis of Parkinson’s disease was made under the age of 50, 100% of the monozygotic (from the same egg, or identical) twins both had the condition, while only 10-20% of the dizygotic (from separate eggs) twins both had Parkinson’s disease. The researchers concluded that ‘this pattern strongly supports a primarily inherited cause of early-onset Parkinson’s disease’.
Curiously, when diagnosis was made over the age of 50 years of age, only 10% of the twin pairs both had Parkinson’s disease (for both monozygotic and dizygotic twins). The Gernsheimer twins are a good example of this (Click here to read more on their case).

Jeff & Jack Gernsheimer in 1982. Source: ReadingEagle
But if identical twins share all of their DNA how could the large differences over 50 years of age be explained?
Some twins may be born with a vulnerability for Parkinson’s disease (like a genetic mutation), but there are some other factors that may also be influential in the initiation of the disease. And this is where scientists start talking about something called epigenetics (“Epi-“, from the Greek for ‘over’ or ‘above’ and “-Genetics”,…well, you should be able to work that one out).
Epigenetics is the study of changes in an organism that are caused by modifications or variations of gene activity rather than alteration of the genetic code itself. These variations in activity may result from external factors that cause genes to turn on and off.

Source: 2ndActHealth
The omnigenics perspective does not discuss this aspect of further complexity. But it too is likely to have an influence in determining aspects of Parkinson’s disease.
So what does it all mean?
A new theory has been proposed by some scientists at Stanford University that is proposing an alternative to how we can interpret genetics-related variance. It places more of an emphasis on the many peripheral genetic variations rather than the core mutations associated with heritable traits. It is based on the results of genetic analysis studies involving 100s of thousands of people.
Does this mean our understanding of the genetics of Parkinson’s disease is all wrong?
No. At present, what has been proposed regarding omnigenics is just a theory. We now need to test it further and see which parts of it work and what parts need to be altered. It would be interesting, however, for some of the Parkinson’s genetics researchers to go back to their big datasets and start looking at all the genetic variations that are just below the threshold of being considered important (or significant). Specifically, to investigate the cellular functions of some of those genes (are they more active in neurons).
This post will have been boring reading for some regular readers, but in the big picture scheme of things the topic is really interesting. Deeper DNA analysis will give us a better understanding of not only genetic variation but also the complex underlying mechanism of conditions like Parkinson’s disease.
I for one liked the idea and thought it may be of interest.
Back to more applicable stuff in the next post!
The banner for todays post was sourced from Pinsdaddy
Hi Simon
That post was not boring, it was absolutely fascinating! Thank you for making a very complex subject (genetics) so understandable and interesting for the not-very-scientific readers! Please keep up the good work – I, and many other fellow Parkinson’s people, look forward very much to receiving your posts.
Best wishes Ken Howard
>
LikeLike
Hi Ken,
Thanks for your comment. I’m glad someone else thought this was interesting. I’m really fascinated by the height research, and I like the idea of using it as a template for other traits and diseases. I wonder if we should focus on breaking down PD into more specific components (such as tremor) and assess the PD genetic datasets just for that.
Anyways, glad you liked it.
Kind regards,
Simon
LikeLike
Hi Simon,
Well I simply can’t resist replying to you any longer. Like Ken that was far from boring. This strikes me as being multifactorial inheritance. Can you tell me the difference between multifactorial inheritance and omnigenics?
Oh! just thought, as other people I have met with PD also have various disorders in their families, (not entirely coincidental on the face of it), do you think some of the other genes involved may be close to the main PD genes and therefore more likely to be inherited together (linked)? Hope that makes sense?
Hilary
LikeLike
Hi Hilary,
Thanks for your message. Glad you liked the post. Multifactorial inheritance deals with both genetics and environment, while omnigenics is only looking at the genes. I would think that in quite a few cases, Parkinson’s disease is most certainly multifactorial. A strong or weak genetic predisposition in addition to exposure to some environmental factor/agent (infection, stress, toxin, etc). While omnigenics may provide us with a novel way of looking at genetics, it won’t give us the full picture for complex conditions. Fascinating idea though.
Regarding other inherited conditions, there has recently been a research report published highlighting genetic associations between PD and various autoimmune conditions (https://www.ncbi.nlm.nih.gov/pubmed/28586827). And there is also the melanoma connection with PD (https://scienceofparkinsons.com/2017/03/03/the-red-headed-mice-of-boston/). So yes, I suspect that the presence of other conditions is not co-incidental. Glad other people don’t find this stuff boring!
Kind regards,
Simon
LikeLike